Key takeaways:
~ We humans have lots of variation in the genes that protect against viral infections.
~ Genetic variants can make you more susceptible to certain pathogens while protecting you from others.
Genetics and Susceptibility to Viral Infections
Our genome is shaped by the pathogens that our ancestors survived. This is pretty cool – you carry specific genetic variants passed down to you from ancestors who lived through diseases and epidemics.
Throughout history, humanity’s biggest threats to survival have been the microscopic pathogens that we now battle successfully with antibiotics, antifungals, vaccines, clean water, etc. All of the genetic variants that gave your ancestors a survival advantage in ages past are still written in your genome today. Pretty cool to think about!
Let me give you a few examples:
- People who carry a sickle-cell anemia mutation are likely to have an ancestor who survived malaria.
- One copy of a cystic fibrosis mutation could have protected your ancestors from dying from a cholera outbreak.
- The CCR5Δ32 variant is protective against HIV / AIDS. It is theorized that it may also have protected against the black death in the Middle Ages.
- About 20% of the population carries a variant that prevents them from secreting their blood type – and this also protects them from getting the norovirus and the rotavirus (aka the stomach flu).[ref] Think of this one as a superpower that you would want to have if stuck aboard a cruise ship where everyone is sick!
How does the body protect against viruses?
Viruses are not considered to be living organisms because they don’t have a cellular structure. Instead, they have to get inside a cellular organism in order to reproduce. Biologists define viruses as small, infectious agents. They can infect your cells and then hijack your own cellular processes to replicate themselves. After replication, they spread to other cells, causing more replication and cell death.
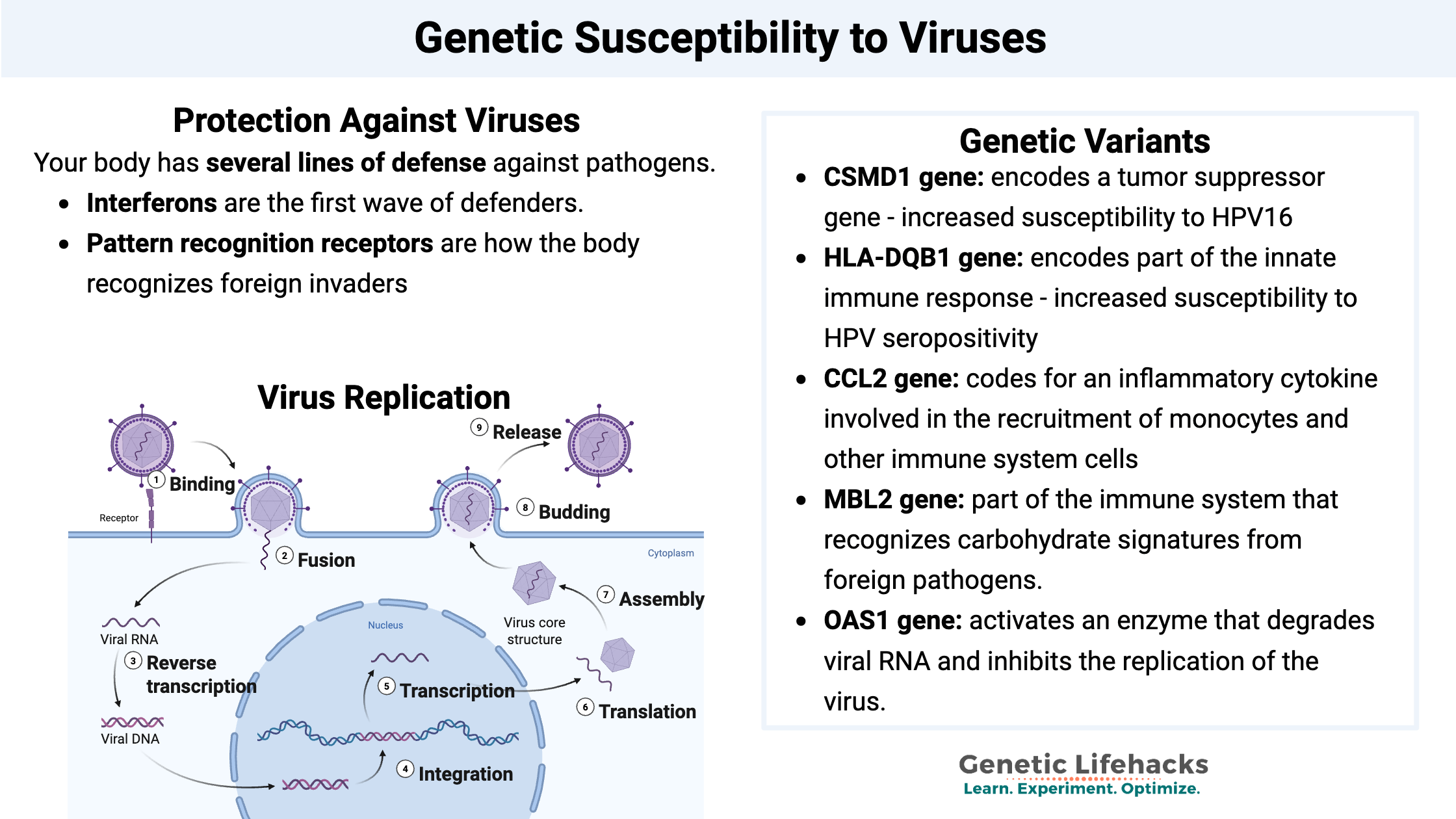
Your body has several lines of defense against pathogens – multiple ways to go to battle, defending the body from out-of-control viral or bacterial invaders. Just like the military has multiple branches (Army, Navy, Air Force, Marines) and specialized groups within those branches, your immune system has several ways of detecting, isolating, killing, and defending against pathogens.
First, the virus must enter the cell. Some viruses need a cell entry receptor to bind to to get into the cell. This is the case with HIV. A mutation in the cell entry receptor CCR5 can prevent HIV from entering a cell and replicating.
Innate immune response:
Even if you’ve never been exposed to a specific virus before, your body has ways to fight it off through your innate immune response.
For viral infections, interferons are the first wave of defenders. White blood cells produce several different types of interferons to act against a wide range of viruses. Interferons act in a couple of different ways:
- Interferons are cytokines that signal to other cells to protect against the spread of the virus.
- Interferons bind to interferon receptors in adjacent cells, which trigger changes in those adjacent cells to resist the viral infection. This basically produces a firewall around the cell that contains the viruses.
- Interferons also stimulate the cells to produce antiviral enzymes.
Pattern recognition receptors are also a part of the innate immune system, which is how the body recognizes foreign invaders. There are several subtypes of pattern recognition receptors, including Toll-like receptors (TLRs), nucleotide-binding oligomerization domain (NOD), retinoic acid-inducible gene 1 (RIG-1) -like receptors, and the C-type lectin receptors (CLRs).[ref]
These pattern recognition receptors are found on immune cells, such as macrophages and neutrophils, and they detect specific parts of a microbe (bacteria, viruses, etc.) and alert the immune system to attack it.
Two specific pattern recognition receptors are TLR3 and TLR4 (toll-like receptor 3/4), and these are what the body uses to identify the SARS coronavirus.[ref][ref]
The HLA (human leukocyte antigen) genes code for part of what is called the major histocompatibility complex (MHC). These proteins are part of the adaptive immune system and present the antigens, or pieces of the virus, on the surface of a cell. These antigens are like a flag saying, ‘the virus is here!’. There are two major divisions of the MHC – MHC class I and MHC class II. The class I antigens that are presented on the surface of the cell call in killer T-cells to destroy the cell. Likewise, the class II antigens present their antigens to the T-lymphocytes, which stimulate antibodies.
How do different genetic variants protect against pathogens?
Genes are the blueprint for the proteins that make up the various components of the immune system. You have genes that code for the proteins that act as cellular receptors, the different pattern recognition receptors, several types of interferon, the various cytokines needed to destroy the pathogens, cell signals, and more. It’s a complicated system that keeps us resilient and able to fight off various types of pathogens, including new viruses and bacteria.
Genetic variants can cause slight differences in how any single part of the immune system works. Each of us is unique – able to easily fight off certain foreign invaders and slightly more susceptible to others. As a whole, this makes the human population resilient and powerful. Some people may be more susceptible to certain pathogens, while others may have a mutation that protects them against that specific foreign invader.
There are, of course, trade-offs. Variants that give protection against a specific pathogen often have a downside, such as increased cancer risk or increased risk of inflammatory conditions[ref][ref]. So if you are battling with an overactive immune response, just keep in mind that it may be the same genetic variant that helped an ancient ancestor to survive a leprosy outbreak and then reproduce, passing along the variant.
This genetic uniqueness comes into play for all of the different aspects of the immune system – from the initial response by interferons to the toll-like receptors and then the HLA proteins that present the antigens on the surface of the cells.
The HLA system comes into play here, big time. There are more than 15,000 different HLA class I and class II alleles that have been identified by researchers so far. There is huge diversity amongst individuals in their HLA types — and this is what gives the human population as a whole a way to survive new pathogens. Because of the dizzying array of different HLA types, it is likely that a portion of the population will be able to fight off novel viruses that have just evolved.[ref]
Viral Susceptibility Genotype Report:
Access this content:
An active subscription is required to access this content.
Lifehacks for preventing viral illnesses
The first steps in preventing contagious diseases are to wash your hands frequently, have good bathroom hygiene, and sleeping well each night. I mentioned above that antibiotics, antifungals, etc., have been important in fighting off pathogens, but really – mankind took a huge step forward in resisting diseases with the implementation of basic hygiene, including simple hand washing (with soap) and having clean water.
Why does handwashing work so well for enveloped viruses such as the coronavirus? Enveloped viruses contain a lipid membrane around the virus, and that lipid (fat) is susceptible to the surfactant actions of soap. Studies on other enveloped viruses, such as the cytomegalovirus, show that good handwashing with soap and water was as effective as hand sanitizer. Other studies, though, show that you may be a little better off with an iodine-based hand sanitizer for Ebola.[ref][ref]
What else may work, according to research, for boosting your ability to fight off a virus?
Access this content:
An active subscription is required to access this content.
Related Articles and Topics:
Familial Mediterranean Fever: MEFV Gene Variants, Overlapping Conditions, and Solutions
References: