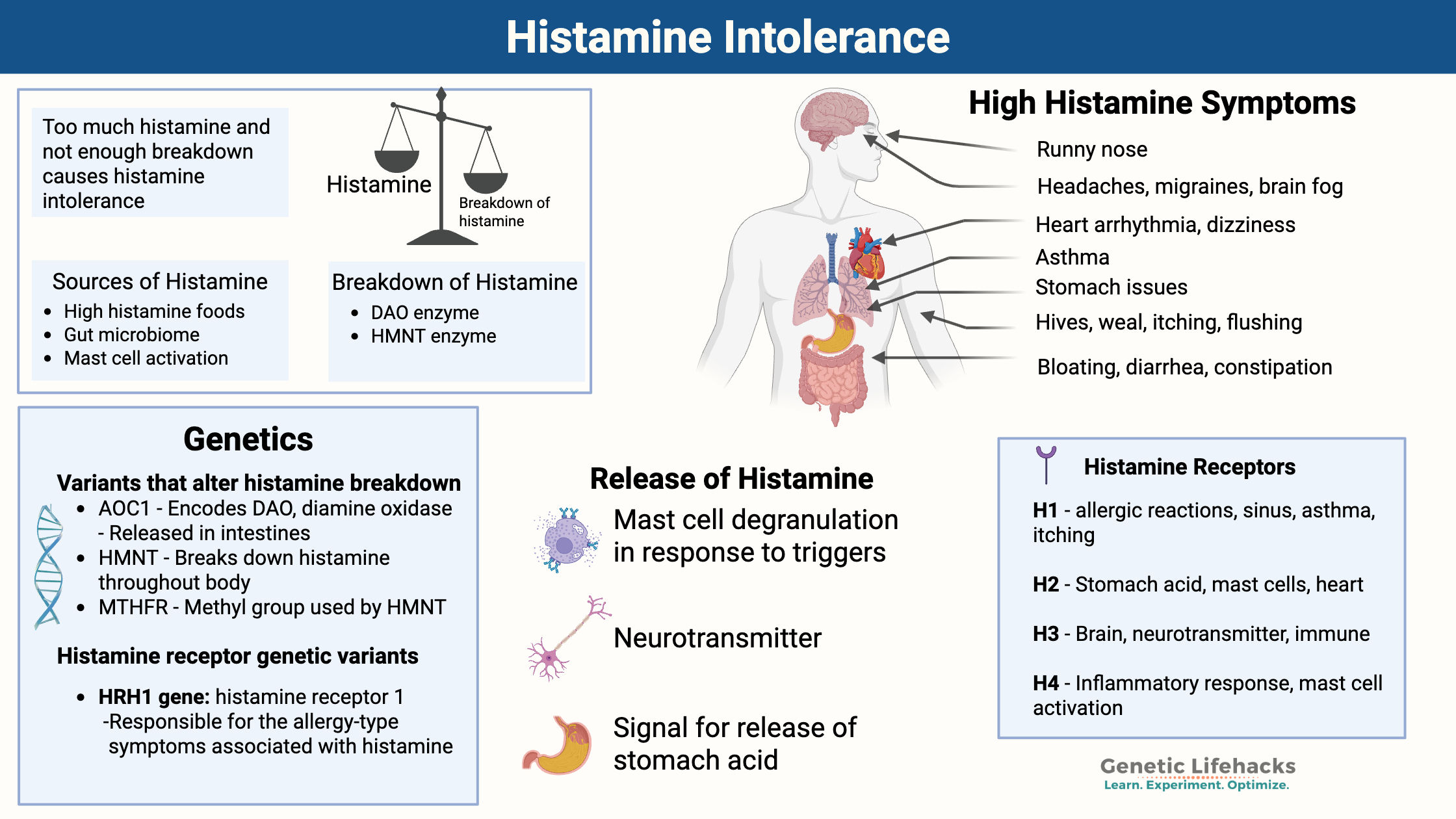
Key takeaways for histamine intolerance:
- Histamine intolerance is caused by an imbalance of too much histamine compared to your capacity to break it down. This can be due to excess histamine from food, producing too much histamine, or not being able to break it down efficiently.
- Genetic variants (SNPs) in the genes encoding DAO and HNMT can reduce enzyme activity, which increases susceptibility to histamine intolerance.
- Histamine intolerance can cause a wide range of symptoms, including headaches, migraines, anxiety, irritability, brain fog, acid reflux, nausea, bloating, diarrhea, arrhythmia, dizziness, sinus drainage, hives, itching, flushing, insomnia, and early waking.
- Understanding your genetic variants can help you target dietary changes, lifestyle modifications, and natural supplements to help with histamine intolerance symptoms.
Want the complete guide? My Histamine Lifehacks book covers the science, connected conditions, supplement protocols by genotype, low-histamine meal plans, and troubleshooting strategies — get a digital copy here for $4.99 Sale. Or buy it on Amazon.
Members will see their genotype report below, plus additional solutions in the Lifehacks section. Consider joining today.
What Causes Histamine Intolerance?
Do you deal with sinus drainage after you eat? Periodic itching and hives? Migraines, irritability, anxiety, and brain fog? The weird and seemingly unrelated symptoms of histamine intolerance can drive you nuts trying to figure out the root cause.
This article is for you if you deal with unexplained headaches, skin reactions, gut problems, or anxiety that seem to come and go — especially if symptoms worsen after certain foods, alcohol, or during allergy season. If you have genetic raw data from 23andMe or AncestryDNA, you can check your histamine-related gene variants below. We will cover the reasons behind histamine intolerance, how your genes influence susceptibility, and end with multiple solutions for eliminating the symptoms.
What is histamine?
Histamine is a biogenic amine that plays many roles in the body. Histamine’s many functions include:
- causes the symptoms in allergic reactions,
- acts within our immune defense system,
- dilates blood vessels (vasodilatation)
- acts as a neurotransmitter
- works as a signaling molecule in the stomach to release acid
While most of us think of histamine in relation to allergies, histamine is a vital part of how your body normally works. The key is that you want histamine in the right amount. Balanced histamine levels are essential.
What are the symptoms of histamine intolerance?
Histamine intolerance results from an imbalance between histamine intake/production and your body’s ability to break it down. Symptoms of high histamine levels are diverse and can affect the head, mood, gut, heart, skin, and sleep.
| Body System | Common Symptoms |
|---|---|
| Head | Headaches, migraines |
| Mood/Brain | Anxiety, irritability, brain fog |
| Stomach | Acid reflux, nausea, stomach pain |
| Intestines | Bloating, diarrhea, constipation |
| Heart | Arrhythmia, dizziness |
| Sinuses | Drainage, congestion |
| Skin | Hives, itching, flushing, dandruff |
| Sleep | Insomnia, early waking |
| Bladder | Urgency, frequency, and leakage |
People with histamine intolerance usually have several of the symptoms above, but likely won’t have all of the symptoms.[ref]
Genetics plays a big role in histamine intolerance. A recent study found that genetic DAO deficiency contributed to histamine intolerance in 79% of patients.[ref] (Members will see their DAO genotypes below in the genotype report.)
Let’s dive into the science of why histamine intolerance occurs. Here’s an overview of what we are going to cover in this article:
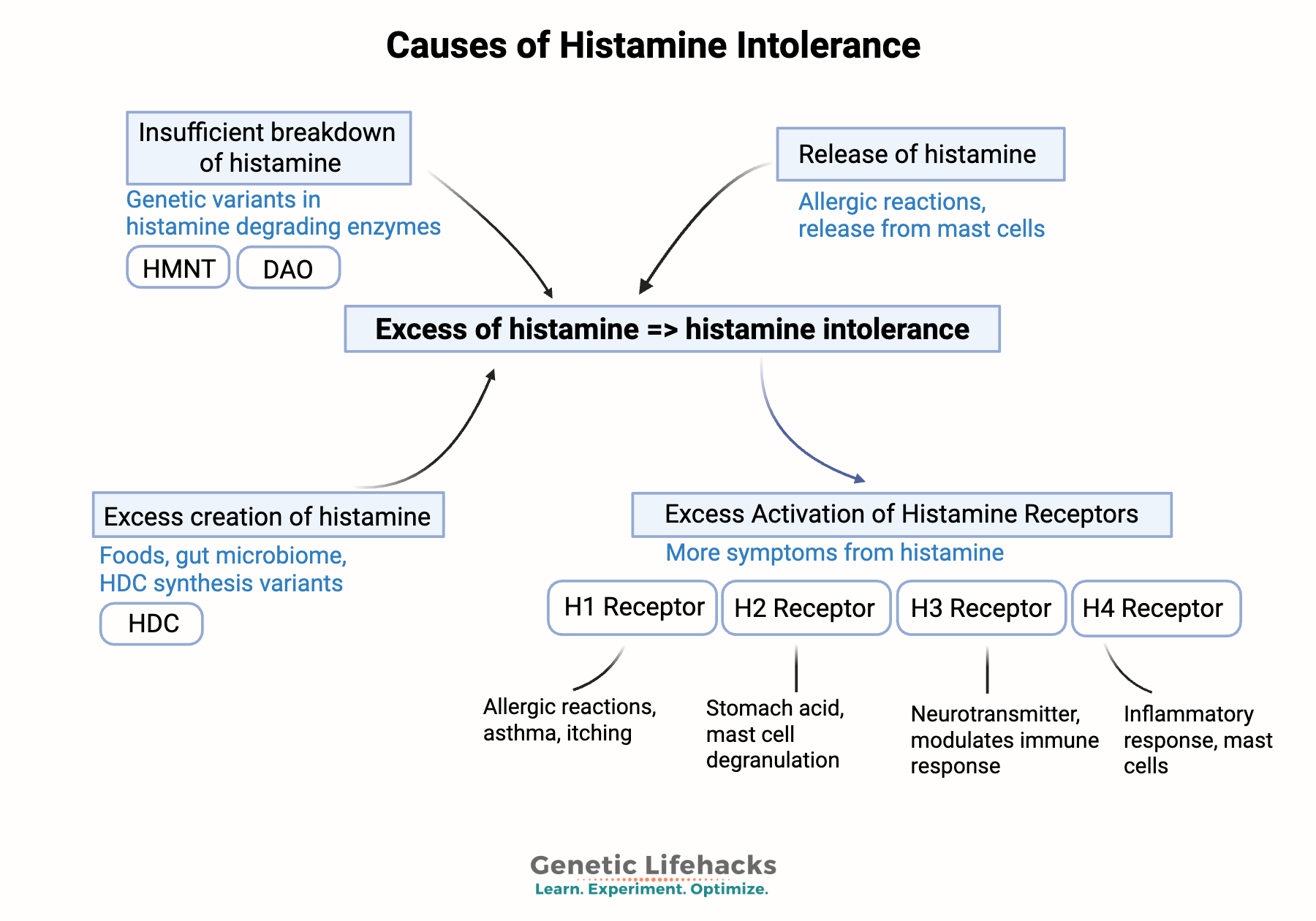
Two Root Causes: Impaired Breakdown vs. Excess Production
There are two main reasons someone might develop histamine intolerance:
- Impaired histamine breakdown due to deficiencies in the DAO and HNMT enzymes.
– and/or- - Excess histamine is being produced, often due to gut dysbiosis, leaky gut, mast cells, HDC variants, or chronic exposure to allergens.
1) Impaired histamine breakdown:
There are two ways your body breaks down and clears histamine: the DAO enzyme or the HMNT enzyme.[ref]
| Enzyme Name | Gene | Location/Action | Main Role |
|---|---|---|---|
| Diamine Oxidase (DAO) | AOC1 | Primarily in the small intestine | Breaks down histamine from foods and gut bacteria [ref] |
| Histamine N-methyltransferase (HNMT) | HNMT | Throughout the body, especially the brain | Breaks down histamine produced by your cells [ref] |
Diamine oxidase (DAO) enzyme: breaking down and inactivating histamine in the gut
Diamine oxidase is encoded by the AOC1 gene. It is primarily produced in the intestines to break down histamine from foods and histamine created by intestinal bacteria. Some species of bacteria in the gut, including those from some probiotics or fermented foods, can add to your body’s histamine level.
- People with histamine intolerance show altered gut microbiome composition as well as elevated levels of zonulin, which regulates tight junctions in the intestines. Excess zonulin causes ‘leaky gut’. [ref]
- A recent study of histamine intolerance patients found that they had “a significantly higher abundance of histamine-secreting bacteria…”[ref]
Histamine plus other biogenic amines: The DAO enzyme is also used by the body to break down other biogenic amines, including tyramine, putrescine, cadaverine, spermidine, and spermine. High levels of other biogenic amines can reduce the ability of DAO to break down histamine.[ref]
DAO degrades histamine into imidazole acetaldehyde, which is then quickly oxidized into imidazole acetic acid.[ref]
Histamine + [DAO enzyme] → Imidazole acetaldehyde + NH3+H2O2
HMNT enzyme inactivates the histamine produced in your cells
In addition to its role in allergic response and in the gut, histamine is produced and functions as a neurotransmitter within the brain and nervous system. The primary enzyme responsible for breaking down histamine in the central nervous system is histamine N-methyltransferase (HNMT). While DAO can also circulate in the periphery, HNMT is the only enzyme that breaks down histamine as a neurotransmitter in the central nervous system and throughout the body.
Genetic variants in HNMT show that brain histamine levels are linked to:[ref][ref][ref]
- Migraines
- ADHD
- Neurodegenerative disorders, such as Parkinson’s disease.
- Cognition
HNMT is active throughout the body, not just in the brain. Genetically decreased HNMT is also linked to:
- Atopic dermatitis or eczema
- Allergies
When HNMT breaks down histamine, it converts it into N-methylhistamine, which is unable to bind to histamine receptors. This reaction uses a methyl group, which ties into the methylation cycle (more on this later). The N-methylhistamine is further broken down with the MAO-B enzyme, forming N-methylimidazole acetaldehyde.[ref]
Histamine (from body cells) → [HNMT enzyme] → N-methylhistamine → [MAO-B enzyme] → N-methylimidazole acetaldehyde
Note that the HNMT enzyme (histamine N-methyltransferase) facilitates the transfer of a methyl group in the breakdown of histamine through this pathway. Thus, having sufficient methyl groups available is important. This ties histamine metabolism into the methylation cycle and MTHFR variants. (more on this in the Genotype report and Lifehacks sections)
2) Excess Histamine Synthesis: The other side of the equation
Histamine is synthesized in cells and then used in multiple ways throughout the body.
Synthesis:
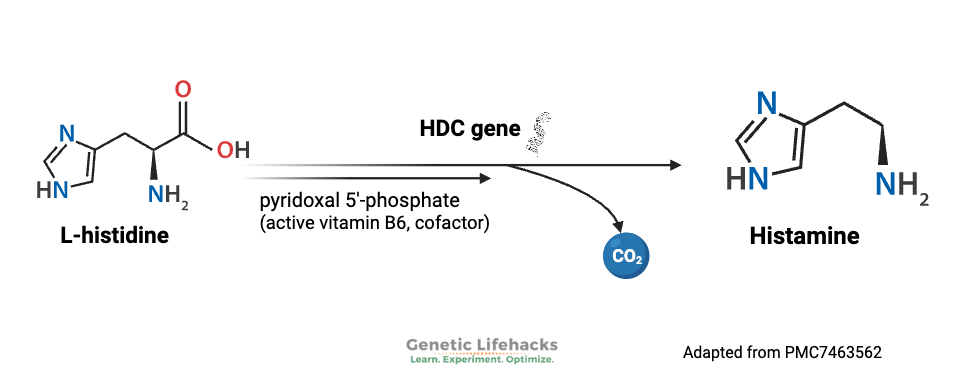
Histamine is made from the amino acid histidine. Histidine is an essential amino acid, meaning humans cannot make it in our bodies and must obtain it from our food. Histidine can be used in the body for several different purposes, including histamine production, metal ion chelation, and neutralizing ROS.
Histidine decarboxylase (HDC gene) is an enzyme that catalyzes the reaction of histidine into histamine. It does this inside various cell types, including creating histamine in large amounts in mast cells.
What happens with too little histamine in the brain?
Without enough histidine decarboxylase (HDC), animals show behavior that resembles Tourette syndrome.
Genetic studies show that people with Tourette’s (vocal and motor tics) can have HDC mutations as a cause. The loss of histamine in the basal ganglia region of the brain causes too much dopamine, resulting in tics.[ref]
Too much histamine affects the heart:
Histamine is also essential in how the heart muscle functions. Too much histamine there can be detrimental, and people with chronic heart failure have higher average plasma histamine levels. In fact, a genetic variant in the HDC gene that reduces histamine levels is linked to a significantly decreased risk of chronic heart failure.[ref]
Additionally, clinical trials show that blocking the H2 receptor is beneficial for chronic heart failure. Famotidine improved cardiac symptoms and ventricular remodeling. In the heart, histamine increases the force of contraction, and even as far back as 1913, histamine has been known to induce heart arrhythmias.[ref][ref]
Mast cells and histamine release:
Mast cells are a type of immune cell that creates and stores histamine. They are found in most tissues in the body, especially in areas of the body exposed to the outside world. Allergens cause mast cells to burst or degranulate and release histamine. Large numbers of mast cells are found in the skin, bronchial tree mucosa, and intestinal mucosa. In addition to allergens, viruses, bacteria, and fungi also activate mast cells.[ref]
Mast cells are also the primary source of histamine released to circulate in the body.[ref]
The release of large amounts of histamine from mast cells can exacerbate problems with histamine from foods. Too many mast cells or mast cells that are too easily activated can lead to mast cell activation syndrome (MCAS). Symptoms of MCAS include flushing, itchy skin, low blood pressure, constipation and/or diarrhea, abdominal pain, shortness of breath, and nasal congestion.[ref] The symptoms of MCAS are driven by excess histamine and overlap with histamine intolerance. Some researchers categorize histamine intolerance as a subset of mast cell activation syndrome (MCAS).
Related articles: Mast cell activation syndrome – Genetics, causes, solutions & MRGPRX2 receptor activation of mast cells
Histamine Receptors: Why Symptoms Vary (H1, H2, H3, H4)
You may wonder why one molecule can cause so many different actions in the body…
People ask, how can histamine cause headaches and heartburn and hives?
The function of histamine in a specific part of the body depends on the receptor it binds to.
Different histamine receptors are found in different parts of the body:[ref]
| Receptor | Location(s) | Main Effects |
|---|---|---|
| H1 | Smooth muscle, CNS, mast cells | Allergy symptoms, vasodilation, asthma |
| H2 | Stomach, blood vessels, heart, CNS | Stomach acid release, heart rhythm |
| H3 | Central/peripheral nervous system (CNS) | Regulates brain histamine and neurotransmitters |
| H4 | Bone marrow, immune cells | Inflammatory response |
H1 receptors: Found in smooth muscle, endothelial cells (lining the blood vessels), the central nervous system, and mast cells. Activating the H1 receptors causes allergy-type symptoms such as itching, swelling, vasodilation, nasal congestion, and skin reactions. H1 receptors are also important in asthma reactions. The H1 receptor is also found in the uterus and plays a role in contractions.
H2 receptors: Acid is released when histamine activates the H2 receptors in the stomach. H2 receptors are also found in the intestinal tract and the walls of blood vessels. Mast cells also have H2 receptors, which, when activated, cause the release of more histamine. In the heart, H2 receptors are essential in controlling the rhythm.
H3 receptors: The central and peripheral nervous systems contain H3 receptors, which act as a feedback loop for histamine levels in the brain. Activating the H3 receptors impacts serotonin, norepinephrine, and acetylcholine release.[ref]
H4 receptors: These histamine receptors are at the core of the inflammatory response. H4 receptors are found in the bone marrow, basophils (a type of white blood cell), the thymus, small intestine, spleen, colon, and mast cells.[ref]
Histamine receptors in the gut:
The H1, H2, and H4 receptors all play a role in the way your intestines function, and this gives rise to the digestive problems seen in people with histamine intolerance.
- Histamine acts as a neurotransmitter involved in the contractions of the intestinal muscles.[ref]
- People with food allergies and IBS had significantly higher levels of H1 and H2 receptors in their intestines.[ref]
Histamine receptors in the brain:
A 2026 study explored the role of histamine and histamine receptors in different regions of the brain. Using single‑nucleus RNA sequencing, the researchers found that H1 and H2 histamine receptors are enriched in excitatory neurons. H3 receptors are preferentially expressed in inhibitory neuronal populations, and HNMT is broadly expressed across neuronal and non‑neuronal cells. The H3 receptors are acting as a feedback loop to restrict histamine and other neurotransmitter release in certain brain regions.[ref]
The study also linked histaminergic neuron expression and changes in the brain to ADHD and major depressive disorder. Altered histamine receptor expression was also seen for anorexia nervosa and schizophrenia.
Genetics and Environment: Combined Effects on Histamine Intolerance
Genetic variants in the genes that encode the histamine-degrading enzymes are strongly linked to an increased relative risk of histamine intolerance. However, research shows that genetics alone is not usually the sole cause. Instead, environmental factors also play a role.
Genetics and Histamine Intolerance:
| Gene | Enzyme/Protein | Impact of Variants | Associated Conditions |
|---|---|---|---|
| AOC1 | DAO (diamine oxidase) | Reduced DAO activity, higher histamine | Migraines, food intolerance |
| HNMT | Histamine N-methyltransferase | Reduced CNS histamine breakdown | ADHD, migraines, eczema, asthma |
| HDC | Histidine decarboxylase | Altered histamine production | Tourette’s, heart conditions |
| MTHFR | Methylene tetrahydrofolate | Reduced methylation | Fewer methyl groups to break down histamine |
AOC1 gene: The AOC1 gene codes for the production of the DAO enzyme. DAO (diamine oxidase) is the enzyme produced in the intestines that breaks down histamine from foods. Genetic variants in AOC1 can increase or decrease the production of the enzyme. These variants are associated with increased risk of histamine intolerance, migraines, and ADHD. [ref][ref]
HNMT gene: Histamine N-methyltransferase (HMNT) is the enzyme that regulates histamine in the body via converting it from histamine into N-methylhistamine, which can then be eliminated from the body. HNMT is responsible for eliminating 80% of histamine in the body.[ref]
HDC gene: The HDC gene encodes an enzyme called histidine decarboxylase, which is used in the conversion of the amino acid histidine into histamine. The enzyme is found primarily in mast cells, neurons, basophils, and cells lining the stomach.[ref] Genetic variants in the HDC gene affect the susceptibility to Tourette’s syndrome and heart disease. [ref]
Methyl groups: The methylation cycle plays a role in breaking down monoamine neurotransmitters, including histamine. It is also important in creating the methyl groups needed for the HMNT enzyme to work. So, looking at your methylation cycle genes can also help with balancing out histamine intolerance.
MTHFR: The MTHFR gene codes for an enzyme that is a key player in the folate cycle. It is one source of methyl groups for the methylation cycle. Decreased enzyme activity of MTHFR, combined with a diet lacking in folate, may result in a reduced breakdown of histamine.
Histamine Receptor Genetic Variants: Histamine is a signal that is relayed by histamine receptors, and genetic variants in the receptors can increase the symptoms by enhancing the signal.
- HRH1 gene: Histamine receptor 1; this receptor is responsible for the allergy-type symptoms associated with histamine (nose running, eyes watering, itchy skin, airway reactivity). Variants in HRH1 affect the risk of allergies.
- HRH2 gene: Histamine receptor 2 is active in the production of stomach acid, the sinus node of the heart, and other places in the body. Variants in HRH2 affect the risk of gastritis, stomach pain, and stomach cancer.
- HRH3 gene: Histamine receptor 3 is important in regulating histamine levels in the brain. Genetic mutations in HRH3 impact the risk of chronic heart failure, and the demyelination of neurons in MS.[ref][ref]
- HRH4 gene: Histamine receptor 4 is found throughout the body, including in the brain. HRH4 receptors are also important in cancer progression.
Environmental Factors Affecting Histamine Intolerance:
For most, histamine intolerance is a combination of genetic susceptibility with environmental factors. By environment, I mean the things that make up your lifestyle – from your sleep to the gut microbiome to the chemicals you’re exposed to every day.
Histamine, Sleep, and Circadian Rhythm:
Histamine is a neurotransmitter in the brain. It is an alerting neurotransmitter, rising in the morning hours to wake us up. About 50% of the histamine in the brain is from mast cells.[ref]
Diphenhydramine (Benadryl) has the side effect of making people sleepy due to blocking the actions of histamine in the brain.
Altering histamine levels in the brain changes sleep:
- In mice, eliminating the histamine receptors in the brain alters sleep patterns. Without histamine, mice were slower to wake up. They also had fragmented sleep and decreased non-REM sleep.[ref]
- In another animal study, researchers decreased the number of mast cells in the brain, reducing histamine production. It did not affect the amount of time the mice slept overall, but it did affect their brain waves in sleep and their ability to bounce back after sleep deprivation.[ref]
In a recent study of people with suspected histamine intolerance, the researchers found that about 25% of the patients had a circadian change in histamine levels that differed from a control group. These patients had significantly reduced DAO enzyme levels and higher histamine levels during the day.[ref]
Your Gut Microbes Produce Histamine:
The gut microbiome significantly influences histamine levels. For some, histamine symptoms begin or worsen when trying to “fix the gut,” especially by consuming fermented foods, which are often high in histamine and may contain histamine-producing bacteria.
Some bacteria carry the HDC gene, which codes for an enzyme that converts L-histidine into histamine. These bacteria may also have transporters that import histidine and export histamine into the gut.[ref]
The gut microbiome in histamine intolerance:
When researchers looked at the gut microbiome, they found significant differences in the gut microbiomes of people with histamine intolerance compared to normal gut microbiomes. Histamine-secreting bacteria were abundant in people with histamine intolerance.[ref] Another study found that people with histamine intolerance found that they had a much lower abundance of Bifidobacteria in the gut and higher levels of Proteobacteria and Roseburia species. The phylum Proteobacteria includes Morganella morganii, which is strongly linked to histamine secretion.[ref]
Environmental chemicals that cause histamine release:
- Sodium fluoride primes mast cells to release histamine. [ref] The addition of sodium fluoride to drinking water is common in most US municipalities.
- PFOAs (Perfluorooctanoic acid) have been found to release histamine and cause mast cell degranulation. “… PFOA exacerbated allergic symptoms via hypothermia, and an increase of serum histamine, TNF-α, IgE, and IgG1 in the ovalbumin-induced systemic anaphylaxis. The present data indicate that PFOA aggravated FcɛRI-mediated mast cell degranulation and allergic symptoms.”[ref] You will find PFOAs in Teflon, stain-resistant carpeting, microwave popcorn bags, food wrappers, etc.
- Off-gassing from carpeting may cause mast cell release in the case of sick building syndrome.[ref]
- Sodium benzoate, a common preservative, causes histamine release in people with allergies and asthma.[ref]
- Aspirin and other salicylates can cause histamine problems for some people, possibly through basophil activation.[ref]
- Polysorbate 80, a common food additive, causes histamine release.[ref] Polysorbate 80 is in most brands of pickles — except organic pickles.
Medications that decrease DAO enzyme production
In addition to foods, drug interactions can cause a decrease in DAO enzyme production.
- Metformin has been shown to decrease the DAO enzyme.[ref]
- Vitamin B3 (nicotinamide or niacinamide) may increase histamine levels at doses of 100 mg or higher.[ref]
Conditions Linked to High Histamine Levels
When discussing histamine intolerance, the conversation often centers on foods and the symptoms affecting the gut or skin. However, elevated histamine levels can contribute to a variety of health conditions. By understanding how diet, histamine-degrading enzymes, the gut microbiome, and the body’s own histamine production are connected, you may be able to help other conditions related to high histamine levels.
Here are other conditions that involve high histamine levels:
- Asthma: Histamine release from mast cells is involved in the airway restriction in asthma.
- Brain fog: High histamine levels can be a cause of brain fog or cognitive issues
- Canker sores, oral lesions, or burning mouth.[ref]
- Periodic Limb Movement Disorder: Animal studies show that H3 receptor activation causes limb movements during sleep
- Autoinflammatory conditions, such as familial Mediterranean fever: People with FMF have higher-than-normal histamine levels during a flare-up.
- Mast Cell Activation Syndrome: MCAS is like histamine intolerance on steroids.
- Ehlers-Danlos Hypermobility Syndrome: Mast cells and high histamine in the joints are a possible underlying cause of hypermobility syndrome.
- Migraines: Genetic variants in the AOC1 (DAO) gene increase the risk of migraines.
- ADHD: Histamine-related variants increase the risk of ADHD
- Early morning waking / insomnia: One underlying cause of early morning insomnia is that high histamine levels cause too much alertness.
- Heart rhythm issues, including atrial fibrillation: The H2 receptor and histamine play an integral role in heart rhythm.
- Fibromyalgia: Mast cell activation and histamine release could be adding to fibromyalgia symptoms.
- COPD: Histamine levels and mast cell activation cause bronchial constriction in COPD.[ref][ref]
- Tinnitus and Meniere’s: A genetic variant in the H4 receptor increases the risk of Meniere’s and ringing in the ears.
- Pollen allergies and food allergies: Mast cells release histamine when triggered by an allergen.
- Chronic fatigue (anecdotal): Many with ME/CFS report problems with mast cell activation or histamine intolerance symptoms.
Note that pregnancy often causes lower histamine levels, and allergies often subside during pregnancy. This is because the placenta produces high levels of the DAO enzyme. This counteracts any activation of H1 receptors in the uterus, which causes contractions.[ref]
How can you use this information?
Histamine intolerance – or rather, overall high histamine levels – could be an additive factor for the above conditions.
If you have a condition that is caused in part by high histamine levels, understanding the dietary and gut microbiome connections to histamine can be a game-changer for some people. For example, if you have brain fog or ringing in your ears, trialing a low-histamine diet for a week or two may help you determine if high-histamine foods are playing a role in your symptoms. Similarly, if you have fibromyalgia, looking at the role of mast cells and histamine may lead to supplements, such as quercetin or DAO, along with a low-histamine diet as a solution.
Genotype Report: Histamine Intolerance
Access this content:
An active subscription is required to access this content.
Lifehacks for histamine intolerance:
Below are the research-backed solutions for histamine intolerance. You may need to try several different ‘lifehacks’ to see which works best for you.
Low-Histamine Diet: What to Eat and What to Avoid
A low-histamine diet restricts foods with high levels of histamine or that cause the body to release histamine. To experiment with a low-histamine diet, eliminate all of the higher-histamine foods for a period of time to see how your body responds.
In general, foods that are fermented or aged are higher in histamine. High histamine foods include processed meats, cheeses (except farmer cheese), fish and seafood that isn’t completely fresh, spinach, chocolate, tomatoes, strawberries, wine, sake, and more.
High histamine foods list:
If you are considering a low histamine diet, I find this histamine food list to be the most thorough:
Complete list of foods that are high in histamine (pdf).
Here are the highlights of what to avoid on a low-histamine diet:
| Raw egg whites | Buckwheat, malted barley | Chickpeas |
| Blue Cheese | Walnuts, pecans | Lentils |
| Hard cheeses (mature) | Avocados | Spinach |
| Dried meat (jerky) | Eggplant | Soybeans, edamame |
| Cured ham | Hot peppers | Tomatoes |
| Most organ meats | Anything pickled | Bananas (very ripe) |
| Processed meats | Citrus fruits | Chocolate |
| Smoked meat, BBQ | Guava | Kiwi |
| Anchovies | Pineapple | Strawberries |
| Fish (not very fresh) | Nori, algae | Cumin |
| Meat (not fresh, leftovers) | Mustard seeds | Soy sauce |
| Seafood (except ‘frozen at sea’) | Vinegar |
You may find that some of these foods don’t cause histamine reactions, so it is a matter of trial and error to find out what works best for you. For example, soy sauce causes histamine reactions in many people, but it may depend on which species of bacteria is used in the fermentation process. Certain bacteria can reduce the histamine content of soy sauce.[ref]
Note that alcohol can cause mast cells to release histamine. For some, this may cause flushing when drinking. But for people with histamine intolerance, even alcoholic drinks that are lower in histamine may still cause a reaction.[ref]
Related article: Alcohol, mast cells, and histamine
What does a low-histamine diet do?
- Decreasing the amount of histamine you take into your body will lower the overall amount of histamine circulating in your body.
- Research studies show that a low histamine diet helps with chronic urticaria (itchiness, hives), migraines, stomach problems, and asthma.[ref][ref]
Should you maintain a low histamine diet long-term?
Trying a low-histamine diet for a period of time can give you a lot of insight into how histamine affects your body, but it may not be a diet you want to continue long-term. A low-histamine diet restricts many healthy foods that you may enjoy, such as spinach, strawberries, and avocados.
Use a low-histamine diet as a tool to learn which histamine-containing foods bother you the most. It can also be a short-term way of getting histamine responses under control.
Vitamin C:
Numerous studies over the past 5 decades have shown that vitamin C, whether from oral supplements or via IV, can increase DAO synthesis and help to reduce histamine symptoms. Studies show that vitamin C helps with histamine-related conditions such as allergic reactions, motion sickness, and even bronchoconstriction, such as in asthma.[ref][ref][ref]
Related article: Vitamin C and your genes: Absorption, transport, and deficiency
Probiotics: What to take and what to avoid
Probiotic supplements can contain histamine-releasing bacteria, which exacerbate histamine intolerance. If you are taking a probiotic, please be sure to check the label to see which species it contains. Note that some bacteria use vitamin B6 in the reactions to produce histamine, so taking B6 along with a histamine-producing probiotic bacteria exacerbates the problem.[ref]
Histamine-producing strains:
Common probiotic strains known to produce histamine:[ref] [ref][ref]
- Lactobacillus diolivorans
- Lactobacillus reuteri
- Lactobacillus casei
- Lactobacillus saerimneri
- Lactobacillus delbrueckii subsp. bulgaricus
- Streptococcus thermophilus
Histamine-lowering probiotics:
Certain probiotic strains have demonstrated the ability to reduce histamine levels in the gut and may be beneficial.[ref][ref]
- Bifidobacterium infantis
- Bifidobacterium longum
- Lactobacillus rhamnosius
Histamine neutral (does not produce histamine):[ref]
- Lactobacillus lactis
- Lactobacillus plantarum
Low FODMAPs diet: histamine and gut problems
Interestingly, a randomized controlled study for people diagnosed with IBS found that a low FODMAPs diet reduced symptoms and reduced histamine levels.
It could mean that a FODMAPs diet works because IBS is related to histamine intolerance – or – it could mean that the people diagnosed with IBS were really dealing with gut-related histamine symptoms.[ref] Additionally, the low FODMAPs diet may help to decrease intestinal barrier permeability.
A low FODMAPs diet cuts out a lot of high-histamine foods, so it could reduce histamine levels by eating fewer foods high in histamine. On the other hand, a low FODMAPs diet impacts the gut microbiome and histamine-producing bacteria. Animal studies also link IBS to mast cell activation in the colon, so changing the gut microbiome with a FODMAPs diet may also impact colonic mast cells.[ref]
Learn more about what is included in a low FODMAPs diet: Starting a Low FODMAPs diet
Related article: Genetic reasons the FODMAPs diet doesn’t work
Supplements for Histamine Intolerance:
When looking at natural supplements for histamine intolerance, there are two categories:
- supplements that break down histamine
- supplements that block mast cells from releasing excess histamine.
You may find that tackling histamine symptoms with a two-pronged approach is most effective.
Supplements for inactivating histamine:
Diamine Oxidase (DAO) enzyme:
The diamine oxidase enzyme is available as a DAO supplement for people who don’t produce enough of the enzyme. A recent study found that histamine intolerance symptoms improved significantly when taking DAO capsules before meals.[ref]
Tip: If you are looking for DAO supplements at a health food store, they often shelve them in the digestive enzymes section.
Looking for an alternative to expensive DAO supplements? Pea shoots – those first few inches of the pea plant that come up in the spring – are naturally high in the DAO enzyme.[ref] You can easily grow pea shoots at home, which are a tasty addition to a salad.
Related article: DAO Enzyme Supplements and Research Studies
Vitamin B6 (P5P) and copper:
Vitamin B6 and copper are both cofactors in the reactions that degrade histamine.[ref]
Pyridoxal-5′-phosphate is the active form of vitamin B6. Foods high in vitamin B6 include salmon, tuna, eggs, milk, beef, and carrots. (article) Copper-rich foods include chocolate, organ meats, shellfish, and nuts.
Related article: Vitamin B6: Genetics, absorption, and deficiency
Blocking mast cell degranulation:
Mast cells are the largest producers of histamine in the body. Decreasing the release of histamine from mast cells can be helpful to reduce the overall burden of histamine in the body.
Magnesium:
Studies show that magnesium prevents the overactivation of mast cells, decreasing histamine release. If you are low or deficient in magnesium, increasing your intake of magnesium-rich foods or supplementing with magnesium may help to stabilize your mast cells.[ref] Additionally, histamine release from mast cells has been shown to decrease cellular magnesium levels, making it more likely that someone with histamine-related problems would need additional magnesium.[ref]
Related article: Magnesium absorption, transport, and deficiency genes
*Quercetin:
Quercetin has also been shown in studies to inhibit mast cells from degranulating. Mast cells are one way that the body releases histamine.[ref]
Related article: Quercetin: Studies and Genetic Connections
*Fisetin:
Another natural flavonol, fisetin has been shown in cell studies to inhibit mast cells from degranulating.[ref][ref][ref]
Related article: Fisetin as an antioxidant, mast cell stabilizer, and senolytic
*Luteolin:
Luteolin is a natural supplement that has been shown in studies to inhibit histamine release from mast cells.[ref]
Related article: Luteolin as a mast cell inhibitor
Your genotype for COMT rs4680 is —, which means your connected data file indicates COMT function. Some people with slow COMT report irritability or mood changes when taking methyl donors or COMT inhibitors. Read the full article on COMT here.
Gluten sensitivity and histamine intolerance
A new study looked at the correlation between histamine intolerance and gluten intolerance symptoms. It concluded that there was a significant overlap in symptoms and that a low-histamine diet may help people with gluten sensitivity.[ref]
Food preparation methods to reduce histamine levels
How food is prepared makes a big difference in the histamine levels.
A recent study concluded, “Frying and grilling increased histamine levels in foods, whereas boiling had little influence or even decreased them. The boiling method might be helpful to control the effect of histamine in histamine-sensitive or susceptible patients, compared with frying and grilling.”[ref]
Additionally, leftovers kept in the fridge (especially meats!) can build up histamine. Instead, try putting your leftovers in the freezer and thawing them when you want to eat them. Histamine levels increase as food starts to break down. Additionally, other biogenic amines increase, which shuttles part of your DAO enzyme production towards breaking down the other biogenic amines.[ref]
Fermenting and drying foods also increases histamine levels.
Reducing histamine levels in food:
Here are a few tips on reducing histamine levels in your foods:
- Avoid leftovers that have been sitting in the fridge for a few days.
- Freeze meat when you bring it home from the grocery store; quickly thaw it on the day you eat it.
- Look for fish or seafood that says “frozen at sea” on the package
- Smoked meats, such as BBQ pork or brisket, can contain very high levels of histamine.
- Foods cooked in a crockpot (low and slow) are more likely to be high in histamine.
Plan ahead:
- If you’re eating out and don’t know how fresh the meat is, take a DAO supplement or quercetin before your meal.
- If you ate pepperoni pizza for dinner (knowing that you’ll regret it :-), take an H2 blocker before bed to prevent heartburn.
- During pollen season, you may find that sticking to a low-histamine diet helps to reduce your allergy symptoms.
OTC medications that may help:
One OTC medication that can help reduce histamine levels is diphenhydramine (Benadryl). Diphenhydramine acts as both an H1 blocker and reduces histamine levels. Most other ‘antihistamine’ medications, such as common allergy medicines, just block the H1 receptor. This may mask some of the symptoms of histamine intolerance, but the high histamine levels remain and symptoms will return immediately when the receptor blocking wears off.
H2 blockers, such as cimetidine and famotidine, may help with histamine-related stomach problems. Again, these OTC medications aren’t something that I personally would want to take daily, but rather something to have on hand when needed.
Recap of your genes:
Access this content:
An active subscription is required to access this content.
Related Food Sensitivities:
FAQ Recap:
What genes cause histamine intolerance?
Genetic variants in the AOC1 gene (encoding the DAO enzyme) and the HNMT gene are the primary genetic factors. DAO breaks down histamine from food in the gut, while HNMT breaks down histamine produced by the body’s own cells, especially in the brain. Variants in the HDC gene and MTHFR gene can also contribute.
Can you test for histamine intolerance genetically?
Yes. If you have raw genetic data from 23andMe or AncestryDNA, you can check for variants in the AOC1, HNMT, and HDC genes that are associated with reduced enzyme activity and increased histamine intolerance risk. Clinical genetic testing for DAO polymorphisms is also available.
What supplements help with histamine intolerance?
DAO enzyme supplements taken before meals help break down dietary histamine. Natural mast cell stabilizers like quercetin, fisetin, and luteolin can reduce histamine release. Vitamin B6 (P5P form) is a cofactor for histamine degradation. Choosing histamine-lowering probiotic strains like Bifidobacterium infantis is also important.
Can histamine intolerance cause anxiety and brain fog?
Yes. Histamine acts as a neurotransmitter in the brain. When HNMT enzyme activity is reduced due to genetic variants, histamine can accumulate in the central nervous system, contributing to anxiety, irritability, brain fog, and sleep disruption.
What is the connection between MCAS and histamine intolerance?
Mast cell activation syndrome (MCAS) involves overactive mast cells releasing excessive histamine. Some researchers categorize histamine intolerance as a subset of MCAS. The symptoms overlap significantly, and both can be influenced by genetic variants in histamine-degrading enzyme genes.
Related Articles and Topics:
References:
Aguilar-Rodea, Pamela, et al. “The Prevalence of Diamine Oxidase Polymorphisms and Their Association with Histamine Intolerance Symptomatology in the Mexican Population.” Biomedicines, vol. 13, no. 9, Sept. 2025, p. 2280. https://doi.org/10.3390/biomedicines13092280
Alm, P. E. “Sodium Fluoride Evoked Histamine Release from Mast Cells: A Study of Cyclic AMP Levels and Effects of Catecholamines.” Agents and Actions, vol. 13, no. 2–3, Apr. 1983, pp. 132–37. https://doi.org/10.1007/BF01967316
Arisawa, Tomiyasu, et al. “Association between Common Genetic Variant of HRH2 and Gastric Cancer Risk.” International Journal of Oncology, vol. 41, no. 2, May 2012, pp. 497–503. https://doi.org/10.3892/ijo.2012.1482
Ayuso, Pedro, et al. “Genetic Variability of Human Diamine Oxidase: Occurrence of Three Nonsynonymous Polymorphisms and Study of Their Effect on Serum Enzyme Activity.” Pharmacogenetics and Genomics, vol. 17, no. 9, Sept. 2007, pp. 687–93. https://doi.org/10.1097/FPC.0b013e328012b8e4
Cai, Wen-Ke, et al. “The HRH4 rs11662595 Mutation Is Associated with Histamine H4 Receptor Dysfunction and with Increased Epithelial-to-Mesenchymal Transition Progress in Non-Small Cell Lung Cancer.” Biochimica et Biophysica Acta: Molecular Basis of Disease, vol. 1863, no. 11, Aug. 2017, pp. 2954–63. https://doi.org/10.1016/j.bbadis.2017.08.018
Che, Denis Nchang, et al. “Fisetin Inhibits IL-31 Production in Stimulated Human Mast Cells: Possibilities of Fisetin Being Exploited to Treat Histamine-Independent Pruritus.” Life Sciences, vol. 201, Mar. 2018, pp. 121–29. https://doi.org/10.1016/j.lfs.2018.03.056
Chikahisa, Sachiko, et al. “Histamine from Brain Resident Mast Cells Promotes Wakefulness and Modulates Behavioral States.” PLoS ONE, vol. 8, no. 10, 2013, e78434. https://doi.org/10.1371/journal.pone.0078434
Chung, Bo Young, et al. “Effect of Different Cooking Methods on Histamine Levels in Selected Foods.” Annals of Dermatology, vol. 29, no. 6, Oct. 2017, pp. 706–14. https://doi.org/10.5021/ad.2017.29.6.706
Cleveland Clinic. “Mast Cell Activation Syndrome.” Cleveland Clinic, https://my.clevelandclinic.org/health/diseases/mast-cell-activation-syndrome. Accessed 14 May 2026.
Duelo, Adriana, et al. “Improvement of Histamine Intolerance Symptoms in Pregnant Women with Diamine Oxidase Deficiency: An Exploratory Study.” Journal of Clinical Medicine, vol. 14, no. 13, June 2025, p. 4573. https://doi.org/10.3390/jcm14134573
Ebbehøj, Niels E., et al. “Outbreak of Eczema and Rhinitis in a Group of Office Workers in Greenland.” International Journal of Circumpolar Health, vol. 74, July 2015, p. 27919. https://doi.org/10.3402/ijch.v74.27919
Ercan-Sencicek, A. Gulhan, et al. “L-Histidine Decarboxylase and Tourette’s Syndrome.” The New England Journal of Medicine, vol. 362, no. 20, May 2010, pp. 1901–08. https://doi.org/10.1056/NEJMoa0907006
Garai, G., et al. “Biogenic Amine Production by Lactic Acid Bacteria Isolated from Cider.” Letters in Applied Microbiology, vol. 45, no. 5, Nov. 2007, pp. 473–78. https://doi.org/10.1111/j.1472-765X.2007.02207.x
García-Martín, Elena, et al. “Diamine Oxidase rs10156191 and rs2052129 Variants Are Associated with the Risk for Migraine.” Headache, vol. 55, no. 2, Jan. 2015, pp. 276–86. https://doi.org/10.1111/head.12493
García-Martín, Elena, et al. “Histamine Pharmacogenomics.” Pharmacogenomics, vol. 10, no. 5, May 2009, pp. 867–83. https://doi.org/10.2217/pgs.09.26
Gervasini, G., et al. “Variability of the L-Histidine Decarboxylase Gene in Allergic Rhinitis.” Allergy, vol. 65, no. 12, Dec. 2010, pp. 1576–84. https://doi.org/10.1111/j.1398-9995.2010.02425.x
Gilligan, Gerardo, and María Fernanda Galindez-Costa. “Histamine Intolerance: A Pioneering Report on the Oral Manifestations of a Complex Systemic Disorder.” Oral Diseases, vol. 31, no. 10, Apr. 2025, pp. 2857–64. https://doi.org/10.1111/odi.15363
He, Gong-Hao, et al. “Associations of Polymorphisms in HRH2, HRH3, DAO, and HNMT Genes with Risk of Chronic Heart Failure.” BioMed Research International, vol. 2016, Feb. 2016, p. 1208476. https://doi.org/10.1155/2016/1208476
He, Gong-Hao, et al. “Relation of Polymorphism of the Histidine Decarboxylase Gene to Chronic Heart Failure in Han Chinese.” The American Journal of Cardiology, vol. 115, no. 11, Mar. 2015, pp. 1555–62. https://doi.org/10.1016/j.amjcard.2015.02.062
Healthline. “Foods High in Vitamin B6.” Healthline, https://www.healthline.com/health/vitamin-b6-foods. Accessed 14 May 2026.
Hon, Yuen Yi, et al. “Endogenous Histamine and Cortisol Levels in Subjects with Different Histamine N-Methyltransferase C314T Genotypes: A Pilot Study.” Molecular Diagnosis & Therapy, vol. 10, no. 2, 2006, pp. 109–14. https://doi.org/10.1007/BF03256450
Hsieh, Chia-Wei, et al. “A Disease Marker for Aspirin-Induced Chronic Urticaria.” International Journal of Molecular Sciences, vol. 15, no. 7, July 2014, pp. 12591–603. https://doi.org/10.3390/ijms150712591
Kim, Jiyoong, et al. “Impact of Blockade of Histamine H2 Receptors on Chronic Heart Failure Revealed by Retrospective and Prospective Randomized Studies.” Journal of the American College of Cardiology, vol. 48, no. 7, Sept. 2006, pp. 1378–84. https://doi.org/10.1016/j.jacc.2006.05.069
Lee, Jun-Kyoung, et al. “Association between Perfluorooctanoic Acid Exposure and Degranulation of Mast Cells in Allergic Inflammation.” Journal of Applied Toxicology, vol. 37, no. 5, Sept. 2016, pp. 554–62. https://doi.org/10.1002/jat.3389
Maintz, L., et al. “Association of Single Nucleotide Polymorphisms in the Diamine Oxidase Gene with Diamine Oxidase Serum Activities.” Allergy, vol. 66, no. 7, Apr. 2011, pp. 893–902. https://doi.org/10.1111/j.1398-9995.2011.02548.x
Martins, Daniel, et al. “Mapping Histamine Pathway Networks in the Human Brain across Cognition and Psychiatric Disorders.” Nature Mental Health, vol. 4, no. 5, May 2026, pp. 816–28. https://doi.org/10.1038/s44220-026-00637-1
Martner-Hewes, P. M., et al. “Vitamin B-6 Nutriture and Plasma Diamine Oxidase Activity in Pregnant Hispanic Teenagers.” The American Journal of Clinical Nutrition, vol. 44, no. 6, Dec. 1986, pp. 907–13. https://doi.org/10.1093/ajcn/44.6.907
McIntosh, Keith, et al. “FODMAPs Alter Symptoms and the Metabolome of Patients with IBS: A Randomised Controlled Trial.” Gut, vol. 66, no. 7, Mar. 2016, pp. 1241–51. https://doi.org/10.1136/gutjnl-2015-311339
Monash University. “Starting the Low FODMAP Diet.” Monash FODMAP, https://www.monashfodmap.com/ibs-central/i-have-ibs/starting-the-low-fodmap-diet/. Accessed 14 May 2026.
Park, Hyo-Hyun, et al. “Anti-Inflammatory Activity of Fisetin in Human Mast Cells (HMC-1).” Pharmacological Research, vol. 55, no. 1, Oct. 2006, pp. 31–37. https://doi.org/10.1016/j.phrs.2006.10.002
Park, Hyo-Hyun, et al. “Flavonoids Inhibit Histamine Release and Expression of Proinflammatory Cytokines in Mast Cells.” Archives of Pharmacal Research, vol. 31, no. 10, Oct. 2008, pp. 1303–11. https://doi.org/10.1007/s12272-001-2110-5
Piliponsky, Adrian M., et al. “Mast Cells in Viral, Bacterial, and Fungal Infection Immunity.” International Journal of Molecular Sciences, vol. 20, no. 12, June 2019, p. 2851. https://doi.org/10.3390/ijms20122851
Pinzer, T. C., et al. “Circadian Profiling Reveals Higher Histamine Plasma Levels and Lower Diamine Oxidase Serum Activities in 24% of Patients with Suspected Histamine Intolerance Compared to Food Allergy and Controls.” Allergy, vol. 73, no. 4, 2018, pp. 949–57. https://doi.org/10.1111/all.13361
Qiu, Shidong, et al. “Complement Activation Associated with Polysorbate 80 in Beagle Dogs.” International Immunopharmacology, vol. 15, no. 1, Nov. 2012, pp. 144–49. https://doi.org/10.1016/j.intimp.2012.10.021
Sánchez-Pérez, Sònia, et al. “Intestinal Dysbiosis in Patients with Histamine Intolerance.” Nutrients, vol. 14, no. 9, Apr. 2022, p. 1774. https://doi.org/10.3390/nu14091774
Schaubschläger, W. W., et al. “Release of Mediators from Human Gastric Mucosa and Blood in Adverse Reactions to Benzoate.” International Archives of Allergy and Applied Immunology, vol. 96, no. 2, 1991, pp. 97–101. https://doi.org/10.1159/000235478
Schink, M., et al. “Microbial Patterns in Patients with Histamine Intolerance.” Journal of Physiology and Pharmacology, vol. 69, no. 4, Dec. 2018. https://doi.org/10.26402/jpp.2018.4.09
Schnedl, Wolfgang J., et al. “Diamine Oxidase Supplementation Improves Symptoms in Patients with Histamine Intolerance.” Food Science and Biotechnology, vol. 28, no. 6, May 2019, pp. 1779–84. https://doi.org/10.1007/s10068-019-00627-3
Schnedl, Wolfgang J., et al. “Non-Celiac Gluten Sensitivity: People without Celiac Disease Avoiding Gluten—Is It Due to Histamine Intolerance?” Inflammation Research, vol. 67, no. 4, Nov. 2017, pp. 279–84. https://doi.org/10.1007/s00011-017-1117-4
Schnedl, Wolfgang J., et al. “Evaluation of Symptoms and Symptom Combinations in Histamine Intolerance.” Intestinal Research, vol. 17, no. 3, 2019, pp. 427–33. https://doi.org/10.5217/ir.2018.00152
Schneider, Erich H., et al. “Modulation of Behavior by the Histaminergic System: Lessons from HDC-, H3R- and H4R-Deficient Mice.” Neuroscience and Biobehavioral Reviews, vol. 47, Aug. 2014, pp. 101–21. https://doi.org/10.1016/j.neubiorev.2014.07.020
Shimbori, Chiko, et al. “Gut Bacteria Interact Directly with Colonic Mast Cells in a Humanized Mouse Model of IBS.” Gut Microbes, vol. 14, no. 1, 2022, p. 2105095. https://doi.org/10.1080/19490976.2022.2105095
Simon, Tünde, et al. “Asthma Endophenotypes and Polymorphisms in the Histamine Receptor HRH4 Gene.” International Archives of Allergy and Immunology, vol. 159, no. 2, May 2012, pp. 109–20. https://doi.org/10.1159/000335919
Son, Jong Hoon, et al. “Histamine Intolerance and Urinary Tract Symptoms.” Journal of Clinical Medicine, vol. 12, no. 21, 2023, p. 6870. https://doi.org/10.3390/jcm12216870
Stevenson, Jim, et al. “The Role of Histamine Degradation Gene Polymorphisms in Moderating the Effects of Food Additives on Children’s ADHD Symptoms.” American Journal of Psychiatry, vol. 167, no. 9, 2010, pp. 1108–15. https://doi.org/10.1176/appi.ajp.2010.09101529
Swiss Interest Group Histamine Intolerance (SIGHI). “Food Compatibility List.” Mastzellaktivierung.info, https://www.mastzellaktivierung.info/downloads/foodlist/21_FoodList_EN_alphabetic_withCateg.pdf. Accessed 14 May 2026.
Thangam, E. B., et al. “The Role of Histamine and Histamine Receptors in Mast Cell-Mediated Allergy and Inflammation: The Hunt for New Therapeutic Targets.” Frontiers in Immunology, vol. 9, 2018, p. 1873. https://doi.org/10.3389/fimmu.2018.01873
Thomas, Carissa M., et al. “Histamine Derived from Probiotic Lactobacillus Reuteri Suppresses TNF via Modulation of PKA and ERK Signaling.” PLoS ONE, vol. 7, no. 2, Feb. 2012, e31951. https://doi.org/10.1371/journal.pone.0031951
Threlfell, Sarah, et al. “Histamine H3 Receptors Inhibit Serotonin Release in Substantia Nigra Pars Reticulata.” The Journal of Neuroscience, vol. 24, no. 40, Oct. 2004, pp. 8704–10. https://doi.org/10.1523/JNEUROSCI.2690-04.2004
Tian, Yan-Jie, et al. “Excess Nicotinamide Increases Plasma Serotonin and Histamine Levels.” Sheng Li Xue Bao (Acta Physiologica Sinica), vol. 65, no. 1, Feb. 2013, pp. 33–38.
Weng, Zuyi, et al. “Quercetin Is More Effective than Cromolyn in Blocking Human Mast Cell Cytokine Release and Inhibits Contact Dermatitis and Photosensitivity in Humans.” PLoS ONE, vol. 7, no. 3, Mar. 2012, e33805. https://doi.org/10.1371/journal.pone.0033805
Yamada, Hideto, et al. “Effects of −1018G>A Polymorphism of HRH2 (rs2607474) on the Severity of Gastric Mucosal Atrophy.” Journal of Gastrointestinal and Liver Diseases, vol. 21, no. 2, June 2012, pp. 139–43.
Yee, Sook Wah, et al. “Prediction and Validation of Enzyme and Transporter Off-Targets for Metformin.” Journal of Pharmacokinetics and Pharmacodynamics, vol. 42, no. 5, Sept. 2015, pp. 463–75. https://doi.org/10.1007/s10928-015-9436-y
Yoshikawa, Takeo, et al. “Histamine N-Methyltransferase in the Brain.” International Journal of Molecular Sciences, vol. 20, no. 3, 2019, p. 737. https://doi.org/10.3390/ijms20030737
Yu, Xiao, et al. “Genetic Lesioning of Histamine Neurons Increases Sleep–Wake Fragmentation and Reveals Their Contribution to Modafinil-Induced Wakefulness.” Sleep, vol. 42, no. 5, 2019, zsz031. https://doi.org/10.1093/sleep/zsz031
Zeng, Zhi, et al. “Disruption of Histamine H2 Receptor Slows Heart Failure Progression through Reducing Myocardial Apoptosis and Fibrosis.” Clinical Science, vol. 127, no. 7, Oct. 2014, pp. 435–48. https://doi.org/10.1042/CS20130716
Zimatkin, S. M., and O. V. Anichtchik. “Alcohol-Histamine Interactions.” Alcohol and Alcoholism, vol. 34, no. 2, 1999, pp. 141–47. https://doi.org/10.1093/alcalc/34.2.141
Additional sources referenced in the article (PDFs, web pages):
“Allergy Nutrition: Pea Seedlings as a Dietary Supplement.” AllergyNutrition.com, http://www.allergynutrition.com/wp-content/uploads/2014/06/Pea-seedlings-as-a-supplement.pdf.
Sander, L. E., et al. “Selective Expression of Histamine Receptors H1R, H2R, and H4R, but Not H3R, in the Human Intestinal Tract.” Gut, vol. 55, no. 4, 2006, pp. 498–504. https://doi.org/10.1136/gut.2004.061762
Smolinska, Sylwia, et al. “Bacterial Histamine-Secreting Genes in the Gut Microbiome.” BMC Genomics, vol. 22, 2021. https://doi.org/10.1186/s12864-021-08004-3
Theoharides, Theoharis C., et al. “Luteolin as a Therapeutic Strategy.” Journal of Biological Regulators and Homeostatic Agents, vol. 23, no. 1, 2009, pp. 1–6.