Key takeaways:
~ Histamine intolerance causes symptoms such as headaches, hives, and heartburn due to excess histamine in the body.
~ Methyl groups are needed in one pathway of histamine metabolism.
~ Genetic variants that affect histamine metabolism may be exacerbated by methylation cycle SNPs.
Members will see their genotype report below, plus additional solutions in the Lifehacks section. Consider joining today.
What does histamine do and how does it interact with methylation?
Histamine is a molecule that plays many roles in the body. It is involved in allergic reactions, plays a role in our immune defense system, acts as a vasodilator, and is a neurotransmitter. While most of us think of histamine only when reaching for an anti-histamine during allergy season, it is a vital part of our body’s everyday functions.
Histamine that is out of balance with the body’s ability to break it down can cause symptoms collectively known as histamine intolerance. Histamine intolerance symptoms include headaches, migraines, anxiety/irritability, acid reflux, nausea, arrhythmia, sinus drainage, and more.
Genetics plays a role in how well your body breaks down histamine. Read on to find out how to check your genetic data for genes involved with histamine levels.
What is histamine intolerance?
The leading causes of histamine intolerance are too few enzymes needed to break down histamine and/or too much histamine production. (gut microbes or mast cells). This article digs into the genes involved in the production of the enzymes that break down histamine.
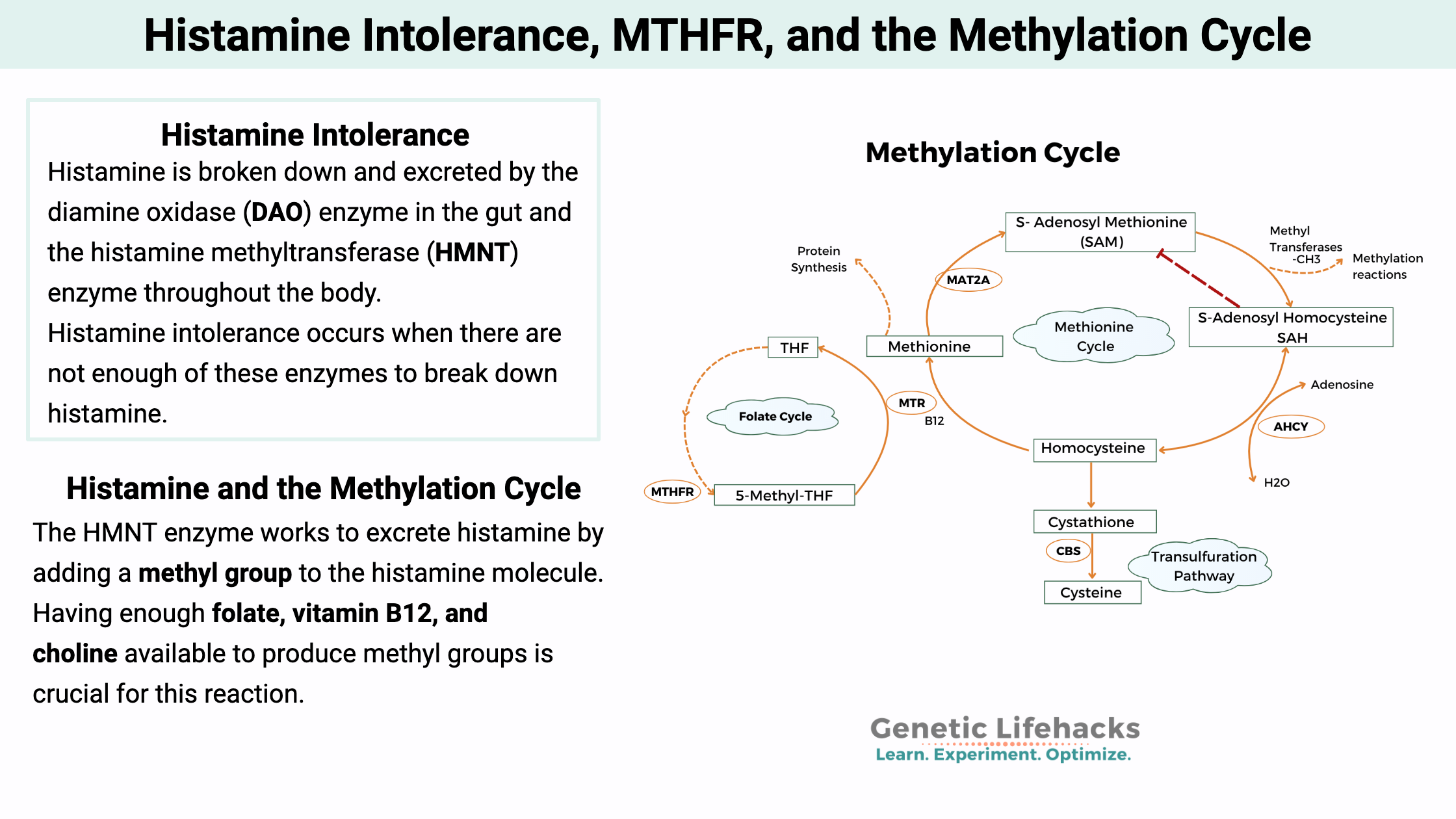
Histamine is broken down and excreted by the diamine oxidase (DAO) enzyme in the gut and the histamine methyltransferase (HMNT) enzyme throughout the body.
Related Article: How your genes increase the risk of histamine intolerance
Histamine and the methylation cycle:
The HNMT enzyme is found throughout the body. This enzyme works to add a methyl group to histamine. The methylated histamine molecules can then be excreted easily in the urine.[ref]
Methyl groups are simply a carbon with three hydrogens. SAMe (s-adenosyl-L-methionine) is the primary donor of methyl groups, and SAMe production relies on the body having enough folate, vitamin B12, and choline available to produce methyl groups.[ref]
Some people have genetic variants that negatively impact the methylation cycle, but fortunately, it is modifiable with a diet high in folate and choline.
Histamine Receptors:
The other side of the histamine equation is the histamine receptors to which histamine binds.
- H1: smooth muscle, endothelium (cells lining the inside of blood vessels and lymph vessels), central nervous system tissue, mast cells (discovered in 1966)
- H2: Gastrointestinal, vascular smooth muscle tissue (walls of blood vessels), mast cells H2 receptors are blocked by cimetidine. (discovered in 1972) “H2 receptors mediate histamine stimulation of gastric acid secretion and may be involved in cardiac stimulation.”
- H3: Central nervous system and some peripheral nervous system, mast cells (discovered in 1987) “feedback inhibitors in CNS.”
- H4: Bone marrow, basophils (a type of white blood cell), thymus, small intestine, spleen, colon, mast cells (discovered in 2001) “considered to have a role in a number of inflammatory disorders such as allergy, asthma, chronic pruritus, and autoimmune diseases.”[ref]
In the intestines, the body’s largest immune organ, three types of histamine receptors have been found: H1, H2, and H4. Low levels of H3 were found in intestinal samples in a few of the study participants. Interestingly, those with food allergies and IBS had significantly higher H1 and H2 receptor levels in their intestines.[ref]
Histamine Genotype Report:
Access this content:
An active subscription is required to access this content.
Lifehacks: Natural Supplements and Dietary Changes
Most people with histamine intolerance find that a low histamine diet can help manage symptoms while getting to the root cause. There are many lists online for foods that are high in histamine or cause the release of histamine. Here is the list that I like to use.
How can you lower histamine levels?
Access this content:
An active subscription is required to access this content.
Related Articles and Genes:
Choline – An Essential Nutrient
An essential nutrient, your need for choline from foods is greatly influenced by your genes. Find out whether you should be adding more choline into your diet.
Quercetin: Scientific studies + genetic connections
Quercetin is a natural flavonoid acting as both an antioxidant and anti-inflammatory. This article focuses on the results of clinical trials involving quercetin as well as linking to specific genetic topics. By using your genetic data, you can make a more informed decision on whether quercetin is worth trying.
Metformin: Longevity research and genetics
A decades-old diabetes drug now holds promise for increasing healthspan. Research shows that metformin may reduce the risk of some of the diseases of aging, thus increasing the number of years someone is healthy.
Folate-rich recipes for MTHFR
Getting enough folate in your diet? We’ve collected some simple nutritional tools and recipes to get you on track.
References:
Ayuso, Pedro, et al. “Genetic Variability of Human Diamine Oxidase: Occurrence of Three Nonsynonymous Polymorphisms and Study of Their Effect on Serum Enzyme Activity.” Pharmacogenetics and Genomics, vol. 17, no. 9, Sept. 2007, pp. 687–93. PubMed, https://doi.org/10.1097/FPC.0b013e328012b8e4.
Chung, Bo Young, et al. “Effect of Different Cooking Methods on Histamine Levels in Selected Foods.” Annals of Dermatology, vol. 29, no. 6, Dec. 2017, pp. 706–14. PubMed, https://doi.org/10.5021/ad.2017.29.6.706.
García-Martín, Elena, Carmen Martínez, et al. “Diamine Oxidase Rs10156191 and Rs2052129 Variants Are Associated with the Risk for Migraine.” Headache, vol. 55, no. 2, Feb. 2015, pp. 276–86. PubMed, https://doi.org/10.1111/head.12493.
García-Martín, Elena, Pedro Ayuso, et al. “Histamine Pharmacogenomics.” Pharmacogenomics, vol. 10, no. 5, May 2009, pp. 867–83. PubMed, https://doi.org/10.2217/pgs.09.26.
Hon, Yuen Yi, et al. “Endogenous Histamine and Cortisol Levels in Subjects with Different Histamine N-Methyltransferase C314T Genotypes : A Pilot Study.” Molecular Diagnosis & Therapy, vol. 10, no. 2, 2006, pp. 109–14. PubMed, https://doi.org/10.1007/BF03256450.
Maintz, L., et al. “Association of Single Nucleotide Polymorphisms in the Diamine Oxidase Gene with Diamine Oxidase Serum Activities.” Allergy, vol. 66, no. 7, July 2011, pp. 893–902. PubMed, https://doi.org/10.1111/j.1398-9995.2011.02548.x.
—. “Association of Single Nucleotide Polymorphisms in the Diamine Oxidase Gene with Diamine Oxidase Serum Activities.” Allergy, vol. 66, no. 7, July 2011, pp. 893–902. PubMed, https://doi.org/10.1111/j.1398-9995.2011.02548.x.
Pang, Y. P., et al. “Theoretical 3D Model of Histamine N-Methyltransferase: Insights into the Effects of a Genetic Polymorphism on Enzymatic Activity and Thermal Stability.” Biochemical and Biophysical Research Communications, vol. 287, no. 1, Sept. 2001, pp. 204–08. PubMed, https://doi.org/10.1006/bbrc.2001.5570.
Sander, L. E., et al. “Selective Expression of Histamine Receptors H1R, H2R, and H4R, but Not H3R, in the Human Intestinal Tract.” Gut, vol. 55, no. 4, Apr. 2006, pp. 498–504. PubMed Central, https://doi.org/10.1136/gut.2004.061762.
Schnedl, Wolfgang J., et al. “Non-Celiac Gluten Sensitivity: People without Celiac Disease Avoiding Gluten-Is It Due to Histamine Intolerance?” Inflammation Research: Official Journal of the European Histamine Research Society … [et Al.], vol. 67, no. 4, Apr. 2018, pp. 279–84. PubMed, https://doi.org/10.1007/s00011-017-1117-4.
Simon, Tünde, et al. “Asthma Endophenotypes and Polymorphisms in the Histamine Receptor HRH4 Gene.” International Archives of Allergy and Immunology, vol. 159, no. 2, 2012, pp. 109–20. PubMed, https://doi.org/10.1159/000335919.
Son, Jee Hee, et al. “A Histamine-Free Diet Is Helpful for Treatment of Adult Patients with Chronic Spontaneous Urticaria.” Annals of Dermatology, vol. 30, no. 2, Apr. 2018, pp. 164–72. PubMed Central, https://doi.org/10.5021/ad.2018.30.2.164.
Weng, Zuyi, et al. “Quercetin Is More Effective than Cromolyn in Blocking Human Mast Cell Cytokine Release and Inhibits Contact Dermatitis and Photosensitivity in Humans.” PloS One, vol. 7, no. 3, 2012, p. e33805. PubMed, https://doi.org/10.1371/journal.pone.0033805.
Yee, Sook Wah, et al. “Prediction and Validation of Enzyme and Transporter Off-Targets for Metformin.” Journal of Pharmacokinetics and Pharmacodynamics, vol. 42, no. 5, Oct. 2015, pp. 463–75. PubMed, https://doi.org/10.1007/s10928-015-9436-y.
Yoshikawa, Takeo, et al. “Histamine N-Methyltransferase in the Brain.” International Journal of Molecular Sciences, vol. 20, no. 3, Feb. 2019, p. 737. PubMed Central, https://doi.org/10.3390/ijms20030737.
—. “Histamine N-Methyltransferase in the Brain.” International Journal of Molecular Sciences, vol. 20, no. 3, Feb. 2019, p. 737. PubMed Central, https://doi.org/10.3390/ijms20030737.