will see their genotype report below and the solutions in the Lifehacks section.
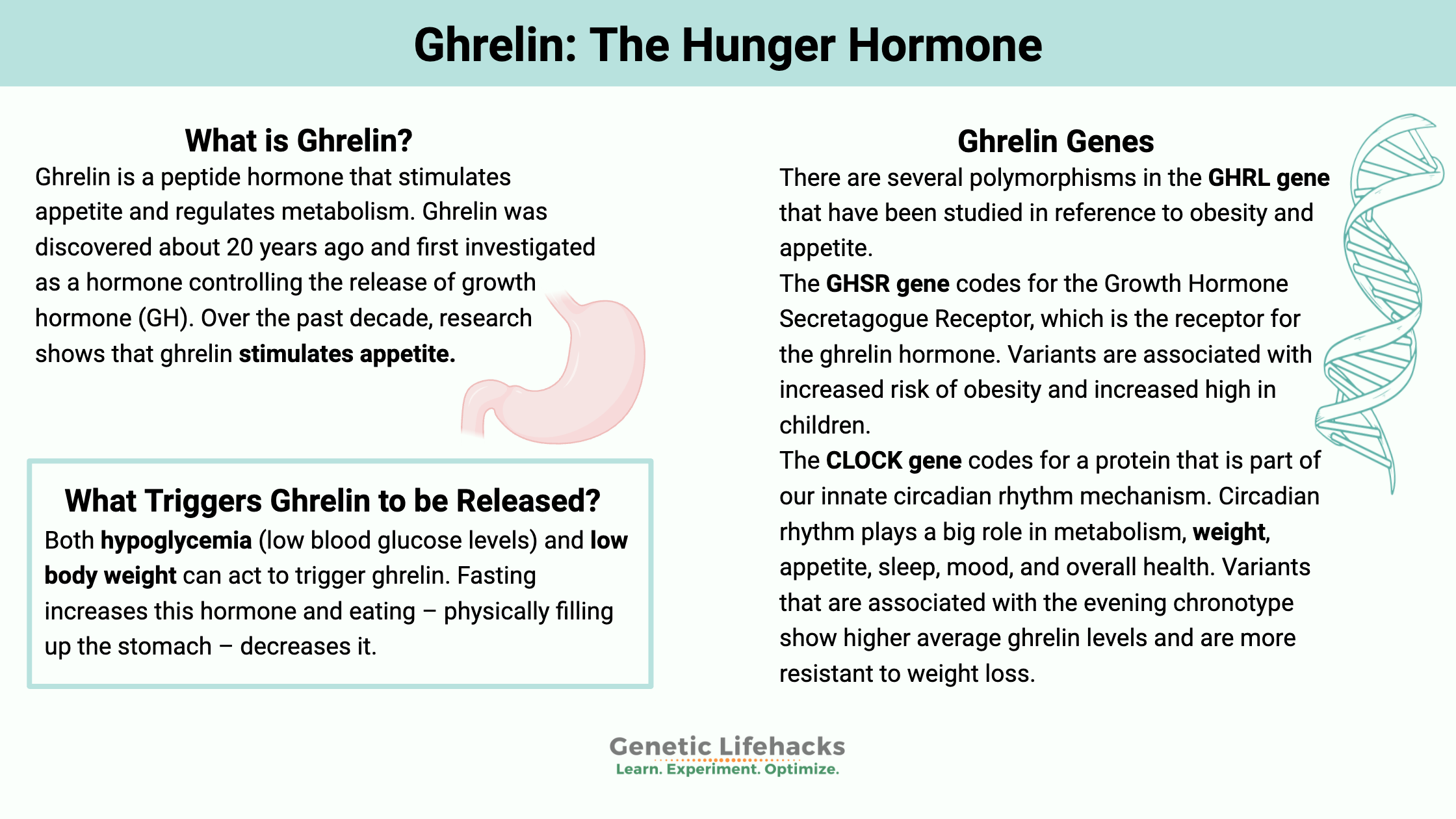
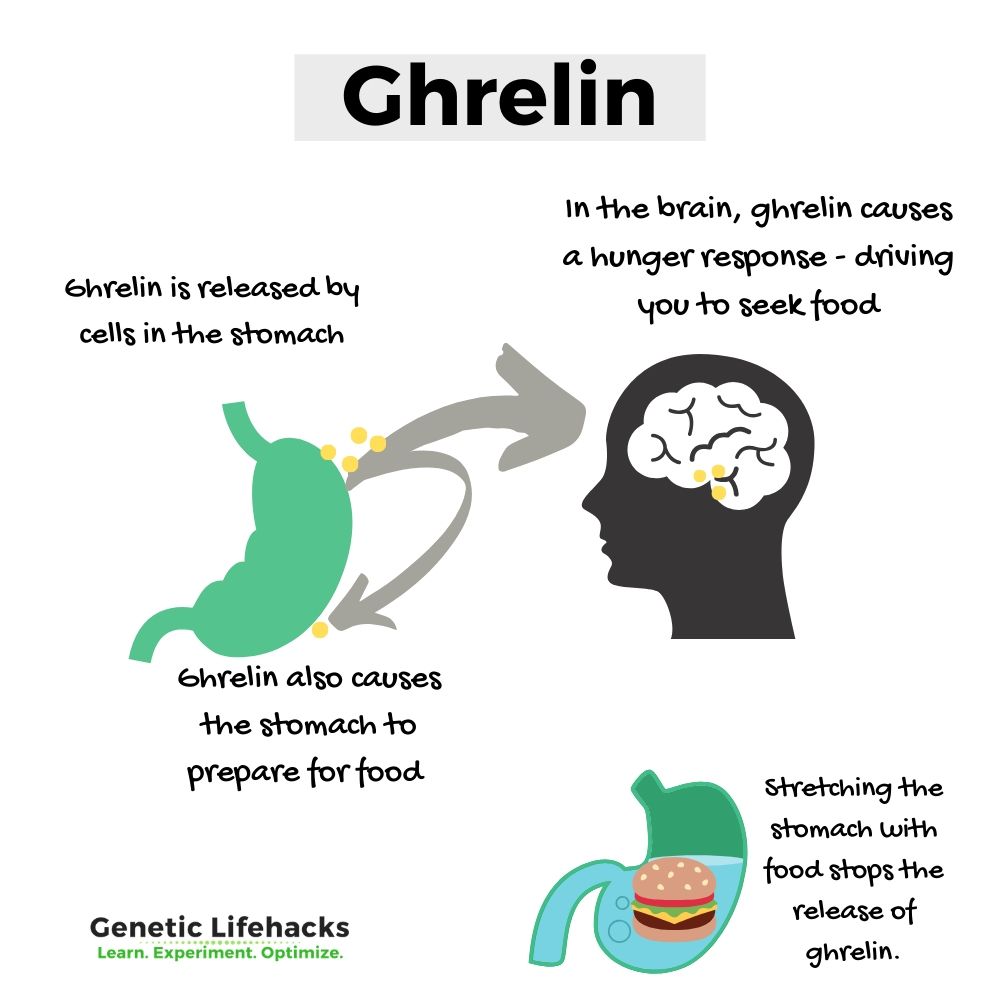
Ghrelin is a peptide hormone that stimulates appetite and regulates metabolism. It drives us to seek out food and eat.
At the most basic level, we are all driven to seek out food. The motivation to eat is inherent to all animals because we simply can’t live without energy from food.
Tasty foods like Doritos and pizza haven’t always been at our fingertips. Our ancestors had to eat to survive. Thus, the innate drive to eat is strong enough to drive people to eat less tasty foods, such as liver, insects, or turnips. This is where ghrelin comes in, prompting us to eat to survive.
Ghrelin was discovered about 20 years ago and first investigated as a hormone controlling the release of growth hormone (GH). Over the past decade, research shows that ghrelin stimulates appetite.[ref]
Thus – the signal sent to the brain prompts you to seek food, and the signal sent to the stomach gets it ready for action.
Fasting increases this hormone and eating – physically filling up the stomach – decreases it.
Researchers don’t have all the answers yet for ghrelin, and there is a lot of ongoing research into the link with BMI, cardiovascular health, and aging.[ref]
Leptin is another hormone that is linked to appetite and weight. It is the counterbalance to ghrelin. Leptin is like the ‘stop’ signal to let you know that you’re full. Just like with ghrelin, there are genetic polymorphisms in the leptin receptor gene that are linked to weight gain.
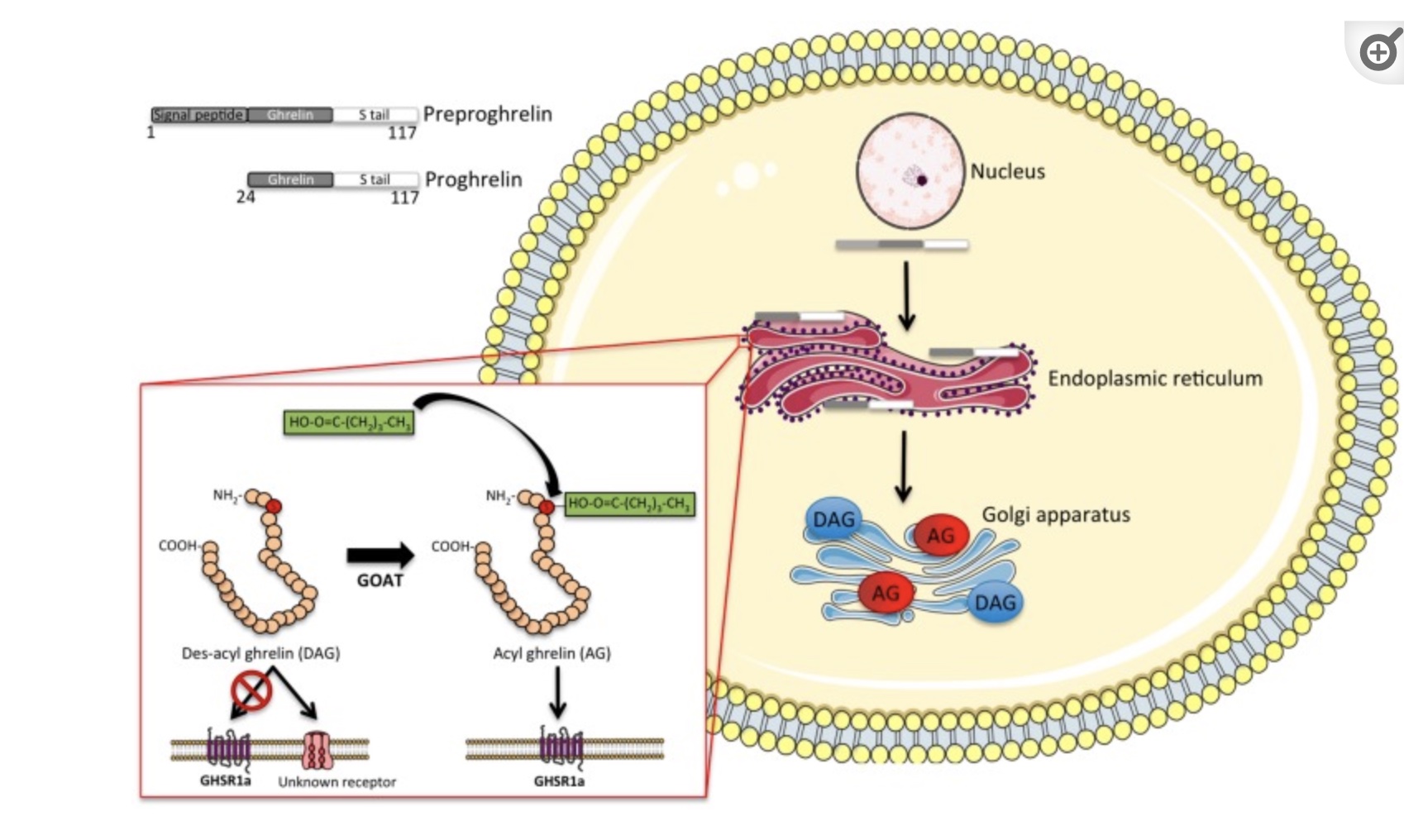
For ghrelin to be in its active form, the enzyme ghrelin-O-acyltransferase (GOAT) acts upon ghrelin to acetylate it. Researchers are investigating drugs that could stop the GOAT enzyme.[ref][ref]
Baessler, Andrea, et al. “Genetic Linkage and Association of the Growth Hormone Secretagogue Receptor (Ghrelin Receptor) Gene in Human Obesity.” Diabetes, vol. 54, no. 1, Jan. 2005, pp. 259–67. PubMed Central, https://doi.org/10.2337/diabetes.54.1.259.
Cabral, Agustina, et al. “Is Ghrelin Synthesized in the Central Nervous System?” International Journal of Molecular Sciences, vol. 18, no. 3, Mar. 2017, p. 638. PubMed Central, https://doi.org/10.3390/ijms18030638.
Claro-Cala, Carmen Maria, et al. “Pomace Olive Oil Concentrated in Triterpenic Acids Restores Vascular Function, Glucose Tolerance and Obesity Progression in Mice.” Nutrients, vol. 12, no. 2, Jan. 2020, p. 323. PubMed, https://doi.org/10.3390/nu12020323.
Gamede, Mlindeli, et al. “The Effects of Plant-Derived Oleanolic Acid on Selected Parameters of Glucose Homeostasis in a Diet-Induced Pre-Diabetic Rat Model.” Molecules (Basel, Switzerland), vol. 23, no. 4, Mar. 2018, p. 794. PubMed, https://doi.org/10.3390/molecules23040794.
Garaulet, Marta, et al. “Ghrelin, Sleep Reduction and Evening Preference: Relationships to CLOCK 3111 T/C SNP and Weight Loss.” PLOS ONE, vol. 6, no. 2, Feb. 2011, p. e17435. PLoS Journals, https://doi.org/10.1371/journal.pone.0017435.
———. “Ghrelin, Sleep Reduction and Evening Preference: Relationships to CLOCK 3111 T/C SNP and Weight Loss.” PLoS ONE, vol. 6, no. 2, Feb. 2011, p. e17435. PubMed Central, https://doi.org/10.1371/journal.pone.0017435.
Gueorguiev, Maria, et al. “Association Studies on Ghrelin and Ghrelin Receptor Gene Polymorphisms with Obesity.” Obesity (Silver Spring, Md.), vol. 17, no. 4, Apr. 2009, pp. 745–54. PubMed, https://doi.org/10.1038/oby.2008.589.
Jakubowicz, Daniela, et al. “Meal Timing and Composition Influence Ghrelin Levels, Appetite Scores and Weight Loss Maintenance in Overweight and Obese Adults.” Steroids, vol. 77, no. 4, Mar. 2012, pp. 323–31. PubMed, https://doi.org/10.1016/j.steroids.2011.12.006.
Kojima, M., et al. “Ghrelin Is a Growth-Hormone-Releasing Acylated Peptide from Stomach.” Nature, vol. 402, no. 6762, Dec. 1999, pp. 656–60. PubMed, https://doi.org/10.1038/45230.
Leeuwendaal, Natasha K., et al. “Gut Peptides and the Microbiome: Focus on Ghrelin.” Current Opinion in Endocrinology, Diabetes, and Obesity, vol. 28, no. 2, Apr. 2021, pp. 243–52. PubMed Central, https://doi.org/10.1097/MED.0000000000000616.
Li, Peng, et al. “Genetic Association Analysis of 30 Genes Related to Obesity in a European American Population.” International Journal of Obesity (2005), vol. 38, no. 5, May 2014, pp. 724–29. PubMed Central, https://doi.org/10.1038/ijo.2013.140.
Mazidi, Mohsen, et al. “The Effect of Hydroalcoholic Extract of Cannabis Sativa on Appetite Hormone in Rat.” Journal of Complementary & Integrative Medicine, vol. 11, no. 4, Dec. 2014, pp. 253–57. PubMed, https://doi.org/10.1515/jcim-2014-0006.
McGavigan, A. K., et al. “L-Cysteine Suppresses Ghrelin and Reduces Appetite in Rodents and Humans.” International Journal of Obesity, vol. 39, no. 3, Mar. 2015, pp. 447–55. www.nature.com, https://doi.org/10.1038/ijo.2014.172.
McGovern-Gooch, Kayleigh R., et al. “Synthetic Triterpenoid Inhibition of Human Ghrelin O-Acyltransferase: Involvement of a Functionally Required Cysteine Provides Mechanistic Insight into Ghrelin Acylation.” Biochemistry, vol. 56, no. 7, Feb. 2017, pp. 919–31. PubMed Central, https://doi.org/10.1021/acs.biochem.6b01008.
Mora, Mireia, et al. “Ghrelin Gene Variants Influence on Metabolic Syndrome Components in Aged Spanish Population.” PLoS ONE, vol. 10, no. 9, Sept. 2015, p. e0136931. PubMed Central, https://doi.org/10.1371/journal.pone.0136931.
Nakajima, Kensuke, et al. “Decreased Plasma Octanoylated Ghrelin Levels in Mice by Oleanolic Acid.” Journal of Oleo Science, vol. 68, no. 1, Jan. 2019, pp. 103–09. PubMed, https://doi.org/10.5650/jos.ess18148.
Parvaresh Rizi, Ehsan, et al. “A High Carbohydrate, but Not Fat or Protein Meal Attenuates Postprandial Ghrelin, PYY and GLP-1 Responses in Chinese Men.” PloS One, vol. 13, no. 1, 2018, p. e0191609. PubMed, https://doi.org/10.1371/journal.pone.0191609.
Perret, Jason, et al. “Polymorphisms for Ghrelin with Consequences on Satiety and Metabolic Alterations.” Current Opinion in Clinical Nutrition and Metabolic Care, vol. 17, no. 4, July 2014, pp. 306–11. PubMed, https://doi.org/10.1097/MCO.0000000000000072.
Riedl, Stefan, et al. “GH Secretagogue Receptor Gene Polymorphisms Are Associated with Stature throughout Childhood.” European Journal of Endocrinology, vol. 166, no. 6, June 2012, pp. 1079–85. PubMed, https://doi.org/10.1530/EJE-11-1112.
Rodriguez-Rodriguez, Rosalia. “Oleanolic Acid and Related Triterpenoids from Olives on Vascular Function: Molecular Mechanisms and Therapeutic Perspectives.” Current Medicinal Chemistry, vol. 22, no. 11, 2015, pp. 1414–25. PubMed, https://doi.org/10.2174/0929867322666141212122921.
Schalla, Martha A., and Andreas Stengel. “Pharmacological Modulation of Ghrelin to Induce Weight Loss: Successes and Challenges.” Current Diabetes Reports, vol. 19, no. 10, Sept. 2019, p. 102. PubMed, https://doi.org/10.1007/s11892-019-1211-9.
Steinle, Nanette I., et al. “Variants in the Ghrelin Gene Are Associated with Metabolic Syndrome in the Old Order Amish.” The Journal of Clinical Endocrinology and Metabolism, vol. 90, no. 12, Dec. 2005, pp. 6672–77. PubMed, https://doi.org/10.1210/jc.2005-0549.
Teuffel, P., et al. “Treatment with the Ghrelin-O-Acyltransferase (GOAT) Inhibitor GO-CoA-Tat Reduces Food Intake by Reducing Meal Frequency in Rats.” Journal of Physiology and Pharmacology: An Official Journal of the Polish Physiological Society, vol. 66, no. 4, Aug. 2015, pp. 493–503.
Yadegari, Mehran, et al. “Interaction between the Genetic Variant of Rs696217-Ghrelin and Food Intake and Obesity and Dyslipidemia.” Annals of Human Genetics, vol. 86, no. 1, Jan. 2022, pp. 14–23. PubMed, https://doi.org/10.1111/ahg.12443.
Yamada, Chihiro, et al. “Ghrelin Enhancer, the Latest Evidence of Rikkunshito.” Frontiers in Nutrition, vol. 8, Dec. 2021, p. 761631. PubMed Central, https://doi.org/10.3389/fnut.2021.761631.
Zhu, Lanjun, et al. “Using the Avocado to Test the Satiety Effects of a Fat-Fiber Combination in Place of Carbohydrate Energy in a Breakfast Meal in Overweight and Obese Men and Women: A Randomized Clinical Trial.” Nutrients, vol. 11, no. 5, Apr. 2019, p. 952. PubMed, https://doi.org/10.3390/nu11050952.