Key takeaways:
~ Many healthy foods contain oxalates, but dietary intake must be balanced with the metabolism and excretion of oxalates.
~ Genetic variants can disrupt the balance of oxalate excretion from the body, leading to the formation of kidney stones or oxalate crystals.
~ Genes also impact the biosynthesis of oxalates in the body.
~ Oxalate crystals can accumulate in the joints, skin, vasculature, and even the brain – leading to inflammation, mitochondrial dysfunction, and cell damage.
Members will see their genotype report below and the solutions in the Lifehacks section. Consider joining today.
What Are Oxalates?
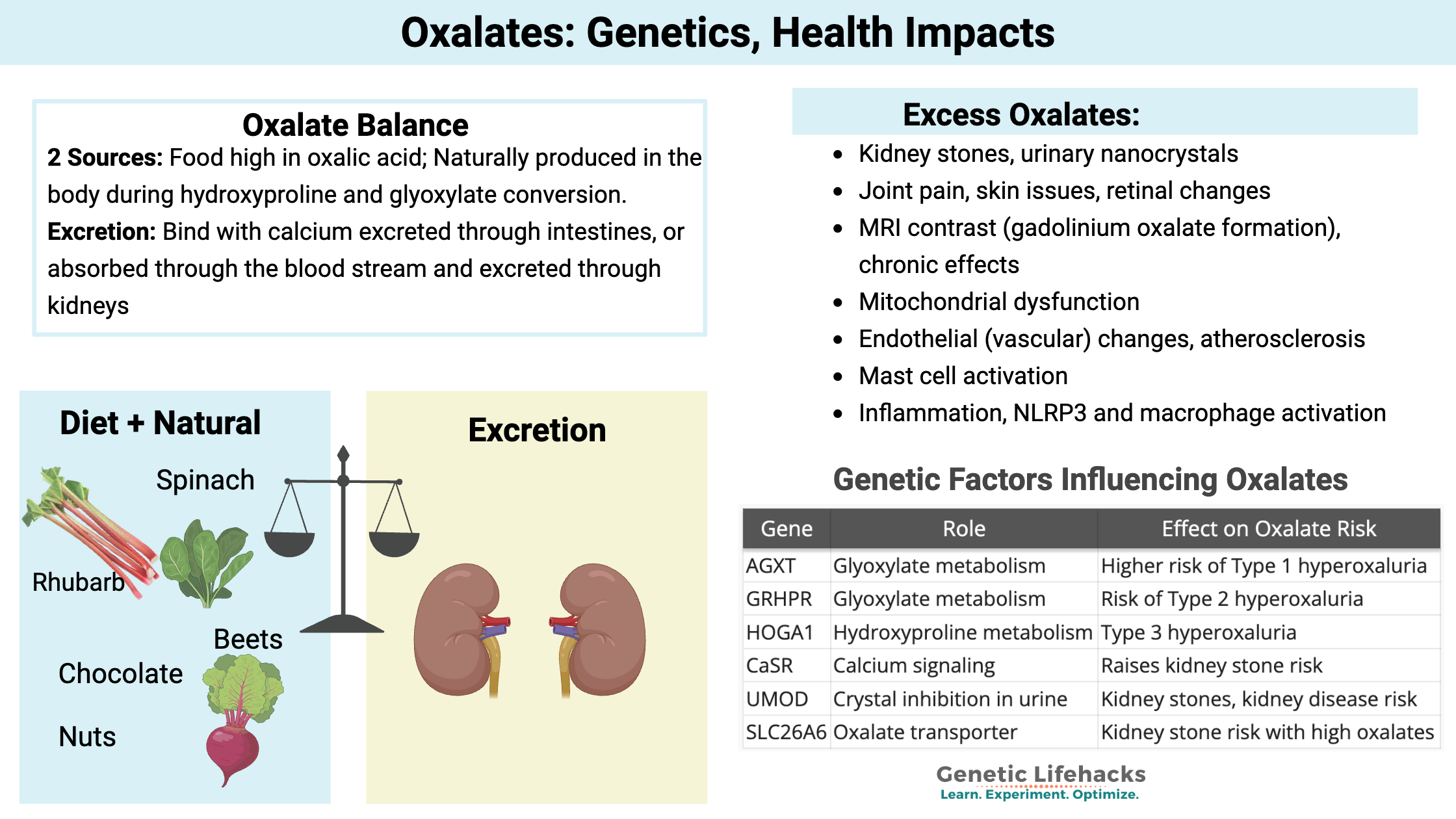
Oxalates are organic compounds found in many of the foods we eat, including healthy greens and delicious chocolate.
We can also produce oxalates in the body through certain pathways. Oxalates can bind with calcium and be excreted through the intestines, or they can enter the bloodstream and eventually be excreted through the kidneys.
Oxalate levels in the body need to be balanced with the breakdown and excretion, matching the production and intake. When oxalate levels are out of balance, problems can arise with oxalate crystal formation, which can cause kidney stones, joint pain, or other problems.
Where do oxalates come from?
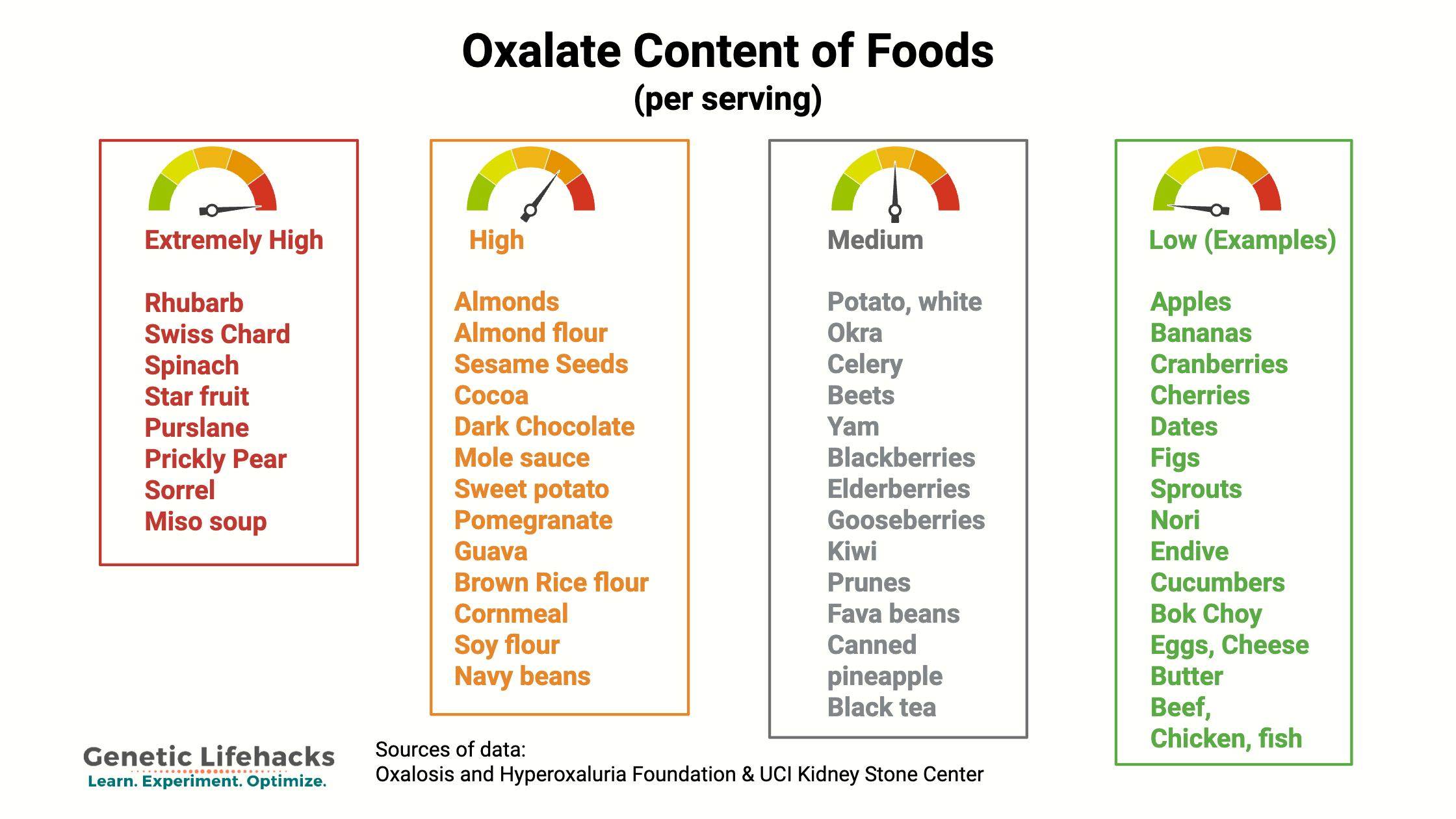
We get oxalates from our diet, especially from foods high in oxalic acid. Oxalates are found in many vegetables and fruits, with some containing high levels of oxalates, while others contain only a tiny amount. Common high-oxalate foods include: Spinach, almonds, beets, rhubarb, sweet potatoes, Swiss chard, and dark chocolate. (Detailed list in the Lifehacks section below.)
When we eat plants high in oxalates, the gut microbiome metabolizes some of the oxalates, and then our intestines absorb some of the oxalates we have eaten. Oxalates are absorbed as free oxalate; however, if calcium is available in the intestines (e.g., from consuming dairy products), it easily binds to oxalates and prevents absorption.[ref]
We can also synthesize oxalate in our bodies through the metabolism of different substances. The biosynthesis of oxalates in the liver accounts for 50-80% of the body’s oxalate pool. Dietary intake accounts for the remainder.[ref]
Precursors of oxalates produced in the body include amino acids and carbohydrate sources. Hydroxyproline, from collagen breakdown, and glycine are amino acid sources that can be metabolized to form oxalates. Glycolate, glyoxylate, and glyoxal are derived from carbohydrates and can form oxalates in the body through their catabolic pathway.[ref]
How the Body Handles Oxalates
When you eat foods that contain oxalates, your gut microbiome plays a big role in how much of the oxalates you will absorb. The bacteria Oxalobacter formigenes uses oxalates as a primary energy source, so the presence of this bacterium can protect against oxalate absorption. One study found that people who had Oxalobacter formigenes in their gut microbiome had lower urinary oxalate levels. The same study found that the absence of Oxalobacter formigenes correlated with higher plasma oxalate levels.[ref]
The kidneys play a key role in maintaining the balance of oxalates in the body. Free oxalates are excreted by the kidneys, or they can combine with calcium ions in the kidneys to form calcium oxalate crystals. If the calcium oxalate crystals infiltrate the vessel walls in the kidneys, this can lead to the formation of kidney stones or even kidney disease.[ref]
Oxalates can also be eliminated in feces when levels are higher in the body. This excretion into the intestine can then be degraded by the gut microbiome and eliminated, or oxalates can be reabsorbed into the body under certain conditions.[ref]
Symptoms of high oxalate levels: Kidney stones, crystals, joint pain, and oxidative stress
Oxalate levels in the body are carefully regulated through a balance of dietary absorption, endogenous production, kidney filtration, intestinal secretion, and reabsorption. When any part of this system is disrupted, oxalate levels can rise abnormally high.
Higher oxalate levels are associated with:
- Kidney stones, urinary nanocrystals
- Joint pain, skin issues, retinal changes
- MRI contrast (gadolinium oxalate formation), chronic effects
- Mitochondrial dysfunction
- Endothelial (vascular) changes, atherosclerosis
- Mast cell activation
- Inflammation and macrophage activation
- NLRP3 inflammasome activation
- Breast cancer (certain types)
Let’s dive into the study details on each of these.
Kidney stones and the urinary tract
High levels of oxalate in the kidneys can lead to the formation of calcium oxalate kidney stones.[ref] Approximately 80% of kidney stones are composed of oxalate bound to calcium. According to a 2005 study, “5% of American women and 12% of men will develop a kidney stone at some time in their life, and prevalence has been rising in both sexes.”[ref]
Urinary crystals can also form in urine that contains calcium and oxalates. These may be the precursors to kidney stones in people with certain changes in their kidney morphology. A study in healthy adults showed that a low-oxalate diet decreased urinary oxalate nanocrystals, and a high-oxalate diet significantly increased urinary oxalate nanocrystals.[ref]
Not everyone with high oxalate levels or with hyperoxaluria will form kidney stones. While we often think of the microbiome in relation to the gut, we also have a urinary tract microbiome. Researchers have found that the diversity and species of bacteria in the kidneys plays a significant role in the formation of kidney stones from oxalate.[ref]
Joint pain, eye problems, skin deposition
In addition to forming kidney stones, oxalate crystals can sometimes be deposited in the joints, skin, and retina. These crystals can deposit in synovial fluid, cartilage, tendons, and bones, triggering an inflammatory response within the joint. Similar to gout, the inflammation causes joint pain, swelling, and stiffness. [ref][ref]
Oxalate-induced arthritis, or joint inflammation due to oxalates, is caused by the deposition of sharp calcium oxalate crystals in the synovial fluid that surrounds a joint. Often, this type of arthritis is seen in people with genetic mutations that cause high oxalate levels.[ref][ref]
Joint inflammation can be either acute, similar to gout, or it can be chronic. Commonly affected joints include ankles, knees, elbows, and knuckles. X-rays sometimes show the calcifications in the joints, in tendons, or in soft tissue near the joint. Oxalate crystal deposition can also cause bursitis and synovitis. [ref]
Tissue remodeling and fibrosis are seen in animal models of hyperoxaluria. This may add to the joint and skin problems seen in people with high oxalate levels.[ref]
Skin issues seen with high oxalates can include papules and nodules on the face and hands, acrocyanosis (bluish hands or feet), or livedo reticularis.[ref]
Mitochondrial dysfunction:
Mitochondria produce ATP (cellular energy molecule) through a pathway involving the inner membrane of this organelle. Oxalate has been shown to depolarize the inner membrane and increase ROS, causing mitochondrial dysfunction and oxidative stress. Other studies show that calcium oxide crystals induce mitochondrial dysfunction in macrophages.[ref][ref]
High dietary oxalate intake causes some, but not all, people to have statistically significant decreases in mitochondrial metabolic function.[ref]
Animal models of high oxalate levels show that there is an increase in mitochondrial reactive oxygen species (ROS) that can lead to oxidative stress if glutathione is inadequate. Mitochondrial dysfunction is found in some people with hyperoxaluria.[ref]
Related article: Glutathione production and genes
MRI Contrast (Gadolinium): Long-term problems
Some individuals have long-term health problems after an MRI with contrast. Kidney damage, joint pain, skin problems, and brain fog top the list. The contrast agent used in some MRIs is gadolinium, which is a metal that enhances the MRI image.[ref]
A 2025 study showed that the presence of oxalic acid causes a decomposition reaction with gadolinium contrast agents (Omniscan and Doterem) to form gadolinium oxalate nanoparticles. Researchers think that the accumulation of gadolinium oxalate nanoparticles in different organ systems may be the cause of the chronic problems some people report after an MRI. It has been known for a decade or more that NLRP3 inflammasome activation occurs after gadolinium-based contrast. [ref][ref][ref]
Endothelial cell damage and atherosclerosis:
Endothelial cells form the inner lining of both blood and lymphatic vessels. A study showed that many types of tissue take up oxalic acid, but in endothelial cells, the oxalate causes a change in calcium levels, leading to endothelial cell injury.[ref]
Endothelial cell injury is directly linked to atherosclerosis. Patients with hyperoxaluria are at a higher risk of vascular dysfunction, including hypertension and atherosclerosis. Some atherosclerotic plaques have been discovered to contain oxalate crystals. A prospective study with an 8-year follow-up showed that higher dietary oxalate levels, especially combined with low calcium, were associated with a higher relative risk of both hypertension and kidney disease. Another study with a 10-year follow-up period found that higher dietary oxalate levels increased the risk of cardiovascular disease incidence by 50%, which was exacerbated by low calcium intake.[ref][ref][ref]
Related article: Cardiovascular disease risk factors
Mast cells:
Mast cells are a type of immune system cell that can quickly degranulate and release histamine, tryptase, and inflammatory cytokines when stimulated by a pathogen, allergen, chemical, or other trigger.
Mast cell activation and histamine intolerance overlap with oxalate sensitivity in a couple of ways:
A study involving cell samples from patients with kidney stones found that they had higher mast cell counts than a healthy control group.[ref] Ketotifen, a drug that stops mast cell degranulation, reduces the formation of calcium oxalate kidney stones.[ref]
Interestingly, the reason that stinging nettles cause severe skin irritation is that they contain oxalic acid and tartaric acid, which have been shown to activate mast cells.[ref][ref]
Related article: Mast cell activation syndrome and Histamine intolerance
Inflammation: Macrophage activation by oxalate crystals
Macrophages are a type of white blood cell that can exist in an anti-inflammatory state (M2) that promotes healing or in a pro-inflammatory state (M1) that increases ROS and inflammation. The pro-inflammatory state (M1) is caused by pathogens, and macrophages are an important way to fight off an infection. However, when the pro-inflammatory state is activated without a pathogen, macrophages can cause damage to the body.
Calcium oxalate crystals are recognized by macrophages as foreign particles, triggering the proinflammatory state and macrophage response.[ref][ref]
A 2021 study looked at the inflammatory response to dietary oxalates. The study participants consumed a low-oxalate diet for 3 days and then consumed a high-oxalate vegetable smoothie. Five hours after the oxalate consumption, the researchers found that crystalline oxalate levels rose, and some individuals had macrophage changes towards inflammation.[ref]
In the liver, oxalates play a role in metabolic dysfunction-associated steatohepatitis (MASH – fatty liver). In people with liver dysfunction, the production of oxalates via lactate dehydrogenase is increased while AGXT (more below on this gene) is decreased. The elevated oxalate levels increase inflammation and fibrosis.[ref]
Related article: Fatty liver disease
NLRP3 inflammasome activation:
The NLRP3 inflammasome is activated by increased ROS and calcium oxalate crystals. NLRP3 then signals for a dramatic increase in inflammatory cytokines. In the case of kidney stones, this leads to inflammation in the kidneys (and pain), which exacerbates the damage to the kidneys. NLRP3 inhibitors are being investigated for kidney stones and kidney disease. Recent studies show that inhibiting NLRP3 activation prevents stone formation by reducing the adhesion of the calcium oxalate crystals in the kidneys.[ref][ref][ref][ref][ref]
Related article: NLRP3 inflammasome and genetic variants
Breast cancer:
The chronic exposure of breast epithelial cells increases the risk that a normal cell will transform into a tumor cell. A study looking at breast tumor tissue found that tumor tissue had higher concentrations of oxalates than comparable non-tumor breast tissue. Microcalcification of calcium oxalate in the breast is associated with invasive carcinomas but also with benign cysts in the breast.[ref]
Sources of Excess Oxalate
High oxalate levels can be due to absorbing excess dietary oxalate, forming excess oxalate in the liver, or not excreting oxalate at the right level through the kidneys.[ref]
| Factor | Mechanism | Risk Group Examples |
|---|---|---|
| Diet | High-oxalate foods | Vegans, raw-food diets |
| Gut microbiome changes | Loss of oxalate-degrading bacteria | Antibiotic users |
| Genetic mutations | Disrupted metabolism | Primary hyperoxaluria |
| Intestinal absorption problems | Increased uptake | Crohn’s, bariatric surgery |
| Kidney function impairment | Poor excretion | Kidney disease patients |
| Vitamin C excess | Conversion to oxalate | Supplement overuse |
| Collagen/gelatin & amino acid breakdown | Precursor conversion | Supplement users, aging |
Let’s look at how each of these can occur in more detail.
1) Oxalate and hydroxyproline in foods:
Foods high in oxalate include spinach, rhubarb, wheat bran, nuts, and chocolate. (See the Lifehacks section for a more complete list.) Consuming a lot of high-oxalate foods, such as a green smoothie made with spinach or lots of dark chocolate, can increase your oxalate levels. For people without additional risk factors, such as genetic variants, kidney disease, or intestinal fat malabsorption, the increased dietary intake is usually excreted through the kidneys without incident.
In addition to oxalic acid in foods, hydroxyproline is an amino acid found in many protein-rich foods that can be metabolized into oxalates. Gelatin and collagen supplements are particularly high in hydroxyproline.[ref]
2) Excess absorption of oxalate:
Intestinal malabsorption problems can increase oxalate levels. Fat malabsorption, whether from exocrine pancreatic insufficiency, bariatric surgery, or IBD, can increase oxalate absorption from the normal 5-10% to over 30%. This 3-6 fold increase in oxalate absorption is significant, especially if paired with a genetic susceptibility or with a high oxalate diet.[ref]
- Enteric hyperoxaluria is a metabolic disorder that is primarily due to increased absorption of dietary oxalates. Typically, this is due to a malabsorption problem from Crohn’s disease, short bowel syndrome, or bariatric surgery.
- Pancreatic enzyme insufficiency can also play a role. Essentially, fatty acids aren’t reabsorbed properly in the small intestine and instead bind to dietary calcium. The absorption of oxalates is regulated in the intestines by being bound to calcium, limiting their absorption, so anything that disrupts calcium in the intestines can affect oxalate levels.[ref]
Related article: Pancreatic Enzyme Insufficiency
3) Endogenous oxalate formation
There are two main sources of oxalate created in the body as a metabolite.[ref]
- Vitamin C (ascorbic acid) is broken down into oxalate without needing enzymes. This means that taking excess vitamin C can increase oxalate levels.
- Glyoxylate is readily converted into oxalate. Glyoxylate is made from glycolate and from the amino acids glycine, serine, and hydroxyproline.
- Glycine can be converted to glyoxylate, but it is usually a minor contributor to oxalate levels.
- Serine can be converted to glycine or it can also be converted to hydroxypyruvate, which can then be converted to glyoxylate with the help of the GRHPR enzyme.
- Hydroxyproline comes from collagen breakdown (or from dietary collagen consumption). It can also be converted to glyoxylate or to glycine.
Hydroxyproline in-depth:
In the kidneys and liver, the amino acid hydroxyproline is metabolized in the mitochondria to form pyruvate (used for ATP production) and glyoxylate.
Hydroxyproline can be catabolized from collagen, such as that found in your connective tissue. The normal daily turnover of collagen in the body is estimated to be 2-3 g/day. [ref]
The AGXT gene encodes the enzyme responsible for breaking down glyoxylate into glycine so that it doesn’t get converted into oxalates.
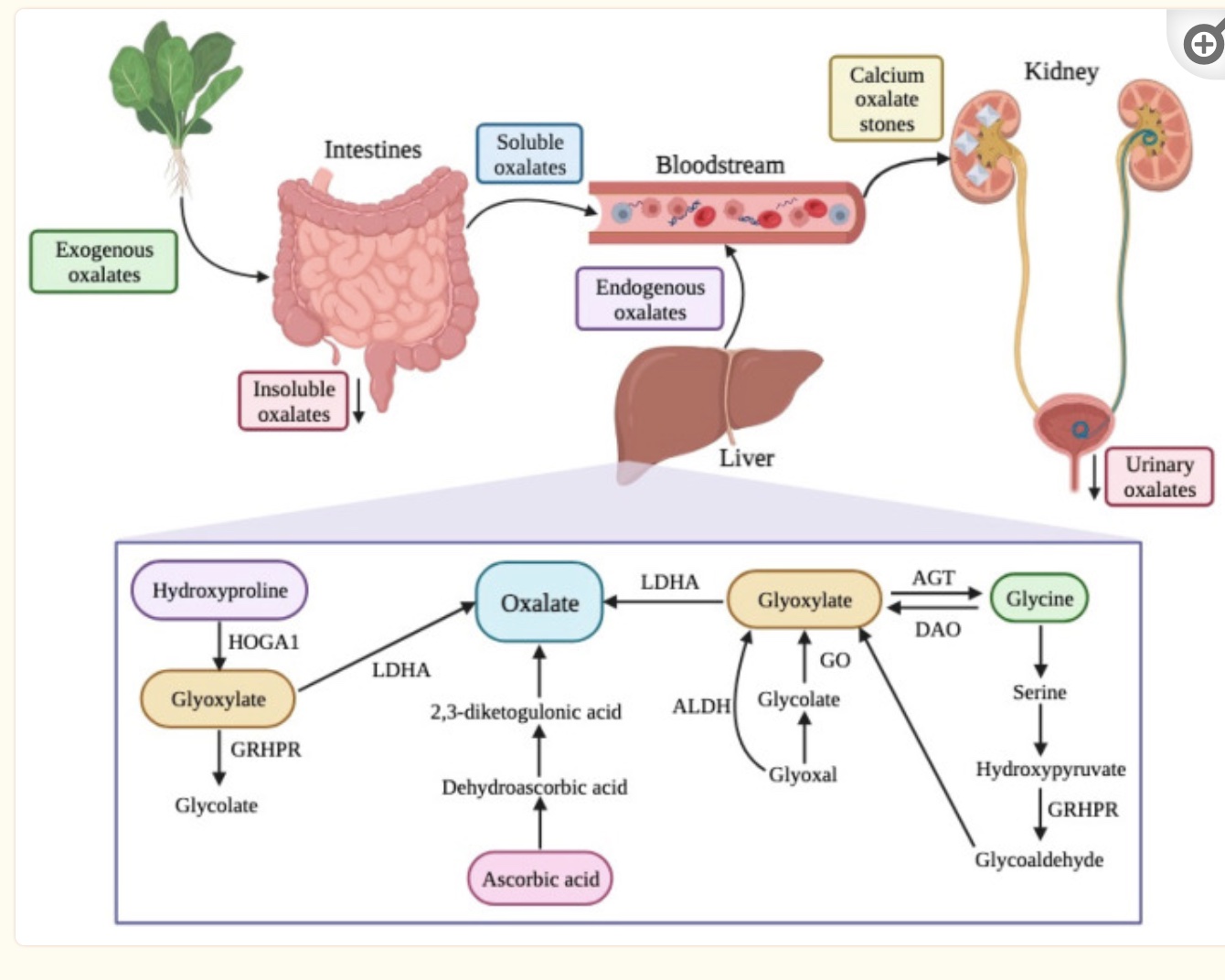
Genetics and Oxalate Metabolism:
Genetic variants in genes related to oxalate formation, transport, and excretion are strongly associated with altered oxalate levels in the body.
Many of the studies on oxalates focus on kidney stone formation, and these studies provide a clear picture of the interaction between oxalate homeostasis, kidney function, and calcium levels. Variants in the CaSR (calcium-sensing receptor) gene increase the risk of kidney stones due to the association between calcium binding to oxalate to form stones.
Primary hyperoxaluria (PH) is caused by genetic mutations that result in high levels of oxalate in the body. People with primary hyperoxaluria are likely to develop kidney stones at a young age. PH can also cause oxalate deposits in most other tissues including the heart, blood vessels, joints, and skin. Bone deformation and osteopathy can also occur.[ref]
There are three types of primary hyperoxaluria: [ref]
- Primary hyperoxaluria type 1 is caused by a mutation in the AGXT gene, which causes a deficiency in the body’s ability to metabolize glyoxylate into other compounds.
- Primary hyperoxaluria type 2 is caused by mutations in the GRHPR gene, which is also involved in glyoxylate metabolism.
- Primary hyperoxaluria type 3 is caused by mutations in the HOGA1 gene, which is involved in the breakdown of hydroxyproline in the mitochondria for use as pyruvate or glyoxalate.
Lactate dehydrogenase (LDHA and LDHB genes) may also act to convert glyoxylate to oxalate in the liver.
Related article: Lactate and lactate dehydrogenase
| Gene | Role | Effect on Oxalate Risk |
|---|---|---|
| AGXT | Glyoxylate metabolism | Higher risk of Type 1 hyperoxaluria |
| GRHPR | Glyoxylate metabolism | Risk of Type 2 hyperoxaluria |
| HOGA1 | Hydroxyproline metabolism | Type 3 hyperoxaluria |
| CaSR | Calcium signaling | Raises kidney stone risk |
| UMOD | Crystal inhibition in urine | Kidney stones, kidney disease risk |
| SLC26A6 | Oxalate transporter | Kidney stone risk with high oxalates |
Let’s dive into these genes in more depth and look at how your genotypes interact with kidney stone risk and higher oxalate levels.
Oxalates Genotype Report
This report section is divided into the following sections:
- Kidney stone risk variants (calcium oxalate stones)
- Primary Hyperoxaluria Type 1 (endogenous oxalate production)
- Primary Hyperoxaluria Type 2
Access this content:
An active subscription is required to access this content.
Lifehacks: Solutions for Oxalate Problems
Please be sure to consult with your healthcare provider if you have any questions on diet, supplements, or your genetic variants. If you have primary hyperoxaluria, as diagnosed by a doctor, be sure to check with your provider about new therapies, such as siRNA therapies that have been recently approved.
Testing your oxalate levels:
An organic acids test, which you can order on your own or through a functional medicine doctor, will show you whether you currently have high levels of oxalate metabolites. Oxalate metabolites include glyceric acid, glycolic acid, and oxalic acid. Glyceric and glycolic acids are elevated in hyperoxaluria.[ref].
You can order an organic acids test through a functional medicine provider — or on your own in most US states (e.g. UltaLabTests).
Here’s a guide to understanding organic acids test results for oxalates. A 2019 study also showed that when just looking at urinary oxalate excretion, levels over 25 mg/day were associated with an increased risk of kidney stones.[ref] Hyperoxaluria is generally defined as urinary oxalate levels >40 mg/day.[ref]
Dietary interventions:
Reducing dietary oxalates or pairing foods containing oxalates with foods that block absorption may help to reduce oxalate crystal formation in the body. In addition, excessive vitamin C intake or collagen intake may also increase oxalates.
Low Oxalate Diet:
If you carry one of the pathogenic hyperoxaluria genetic variants, talk with your doctor and consider whether it would be appropriate to adopt a diet lower in oxalates.
High oxalate foods include:[ref][ref][ref][ref][ref]
Oxalate dumping:
Some people report side effects when suddenly going on a low-oxalate diet. This is attributed to oxalate crystals being released into the bloodstream. There isn’t any peer-reviewed research on this that I could find, but there are a lot of anecdotal reports of oxalate dumping. Thus, reducing oxalate content in your diet more slowly may be a safer way to go.[ref]
Reducing absorption of oxalates from foods:
Access this content:
An active subscription is required to access this content.