Key takeaways:
- The gut microbiome is unique to your body, and it impacts your health in multiple ways.
- Your genes partially decide which microbes can live in your gut, and you can nudge that balance with diet and targeted probiotics.
- A handful of genes have outsized microbiome effects. Variants in genes like FUT2, LCT, VDR, SLC39A8, and APOA5 are each linked to characteristic shifts in specific bacterial groups.
- The Lifehacks section links each gene signal to concrete options such as choosing lactose‑free dairy or lactase‑supportive probiotics, selecting Bifidobacterium‑containing probiotics, adjusting manganese intake, or combining vitamin D with specific Lactobacillus strains.
Your gut microbiome and your genes:
Your gut microbiome is unique to you. But why? Have you ever wondered why our gut microbes are different?
The microbes that take up residence in your intestines depend on:
- the foods you eat (or rather, what you feed your bacteria)
- exposure to different bacteria
- exposure to toxins or antibiotics
- and your genes!
The human gut microbiome is made up of bacteria, viruses, fungi, archaea, and obelisks that inhabit the ecosystem of our intestinal tract. It plays an integral role in our overall health.
Research increasingly shows that host genetics – your gene variants – play an important role in shaping which microbes thrive in our gut. Your genetic variants interact with environmental influences like diet, lifestyle, and medication to shape your gut microbiome.
Why should you care about this? You should care because your genes quietly shape which gut microbes can live in you, and those microbes in turn influence everything from weight and mood to infection risk and cardiovascular health.
How your genes impact your gut microbiome:
Research shows that the influence of our gut microbes on our health is huge.
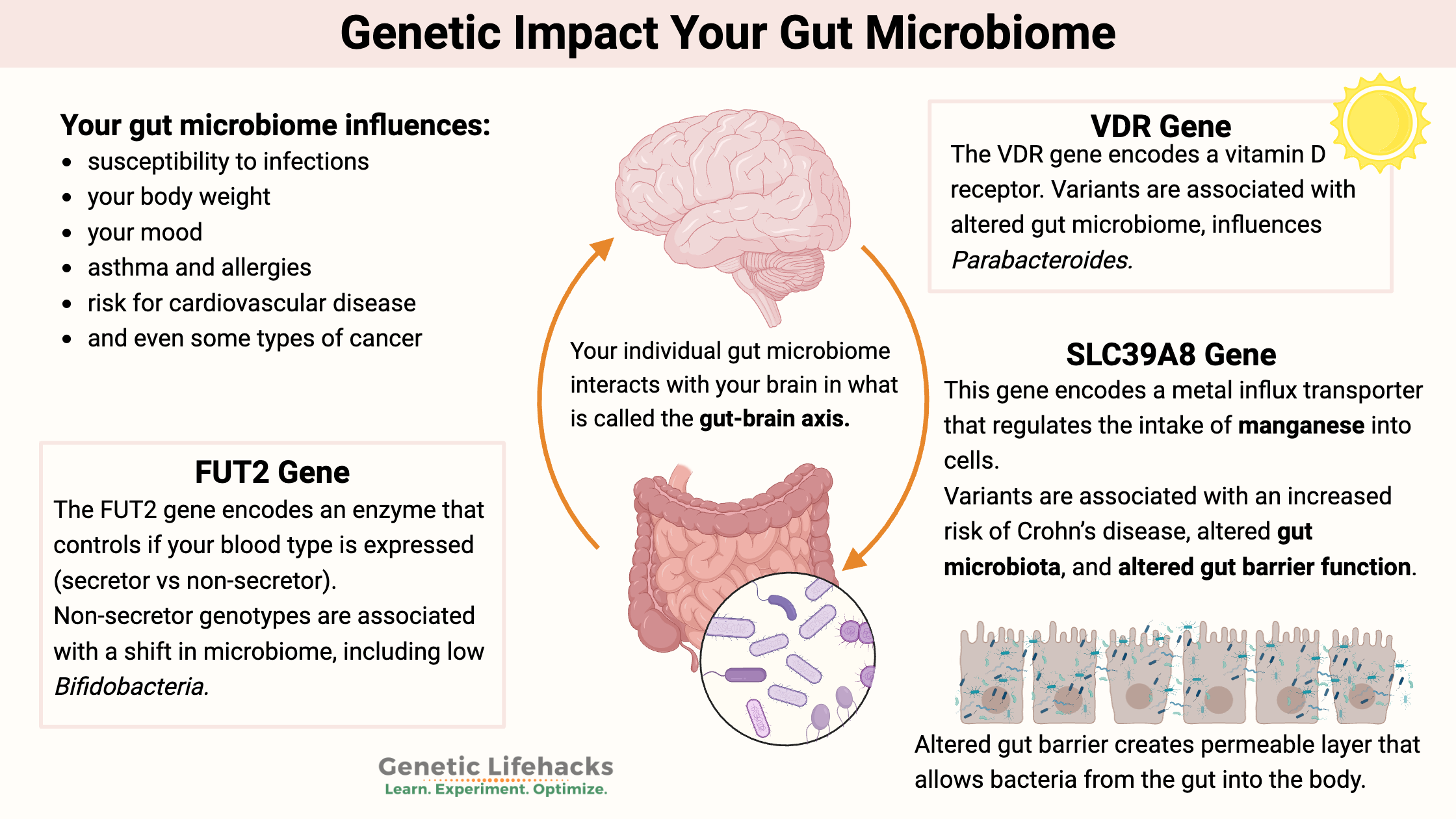
Your gut microbiome influences:
- Susceptibility to infections, such as H. pylori[ref]
- Your body weight (big time)[ref]
- Your mood — including regulating anxiety and depression[ref]
- Whether you have asthma and allergies[ref]
- Your risk for cardiovascular disease[ref]
- Even some types of cancer[ref]
While it may seem odd to think of yourself as a host, as an environment… that is just what you are to the microbes living within or on you. And just like your genetic variants make you look and act differently than everyone else, your unique makeup also influences the types of microbes that can flourish within you.
Heritability and twin studies on the gut microbiome:
Researchers use twin and sibling studies to determine whether something is likely due to heredity (genes + prenatal conditions) or due to being in a shared environment. Nature vs. nurture, to some extent.
Studies of twins and families have revealed that certain components of the gut microbiome are heritable. This means that genetic variants partly explain why some people naturally have higher levels of specific microbes than others. This is especially true for specific species, such as the connection between lactose-loving species and genetic lactose intolerance. But studies also show that living in the same household can also affect the similarity of the microbiome.[ref][ref][ref]
Why is your gut microbiome important?
Whether bloating, rumbling, or running to the bathroom, we have all experienced the misery at some point of not having happy intestinal function… But when it’s working just fine, most people tend to ignore the tiny microbes hard at work in their intestines.
Your gut microbes can play an important role in your health, though.[ref]
- Some microbes produce vitamins such as vitamin K and some B vitamins.
- Others produce short-chain fatty acids used by the intestinal cells as well as signaling to the liver to decrease cholesterol synthesis.
In people who are overweight or obese, the gut microbiome increases how much energy is obtained from food.[ref] You are what you eat — and also what your microbes eat and produce fatty acids from.
Gut Microbes Genotype Report:
Your genetic variants play a role in which species are likely to make up your gut microbiome.
Here we will cover:
- FUT2 helps decide whether you’re a ‘secretor,’ which strongly shapes how many Bifidobacteria and other beneficial microbes can live in your gut, influencing digestion, immunity, and norovirus susceptibility.
- VDR affects how your cells respond to vitamin D and microbial metabolites, which in turn can shift gut diversity and may influence inflammation and conditions like inflammatory bowel disease.
- LCT determines whether you still make lactase as an adult, so it can push your microbiome toward more lactose‑digesting bacteria and change how your gut handles dairy.
- SLC39A8 controls manganese transport into cells, and certain variants are linked to a ‘leakier’ gut barrier and a loss of key short‑chain‑fatty‑acid–producing microbes.
- APOA5 variants can raise triglycerides and are tied to lower Bifidobacteria levels, linking your lipid genetics directly to shifts in your gut microbiome profile.
- ALDH1L1 influences folate metabolism and formate oxidation, and specific variants have been associated with changes in gut microbial metabolites that may impact blood pressure regulation.
- IL4 shapes anti‑inflammatory immune signaling, and certain variants increase susceptibility to C. difficile infection, highlighting how your immune genes affect which pathogens can gain a foothold.
- TLR4 is a sensor for gram‑negative bacteria, and variants here are linked to differences in infection risk, metabolic syndrome, and even H. pylori susceptibility via altered immune–microbiome crosstalk.
Lifehacks:
Work with your gut microbes instead of against them:
Access this content:
An active subscription is required to access this content.
Related Articles and Topics:
TNF-alpha: Inflammation, Chronic Diseases, and Genetic Susceptibility
References: