Primary sclerosing cholangitis (PSC) is a life-altering condition that impacts the liver. For anyone dealing with PSC, research over the past couple of decades points to ways to mitigate some of the damage.
This article was written in conjunction with a Genetic Lifehacks member who has been on a personal quest to understand the PSC research and help out a family member. He wanted to share his collection of research studies with a wider audience to help others struggling with a PSC diagnosis.
Members will see their genotype report below and the solutions in the Lifehacks section. Consider joining today.
What is primary sclerosing cholangitis (PSC)?
Primary sclerosing cholangitis (or PSC) is a rare condition affecting the bile ducts. Your liver produces bile to break down fats into fatty acids that can be easily absorbed from foods. Bile travels from the liver to the gallbladder via the bile ducts.
It is a one-way street – the liver produces bile that travels to the gallbladder and then is released into the intestines.
In PSC, the bile ducts in and outside the liver have been narrowed and scarred. The duct damage then causes the bile to backflow into the liver. As a result, the liver suffers permanent damage.
For people with PSC, this lifelong condition can result in liver damage to the point of often needing a liver transplant.
What causes primary sclerosing cholangitis?
The definite cause of primary sclerosing cholangitis is still not entirely known. It is often diagnosed around 30-40 years of age and is more common in men.[ref]
Essentially, there is an inflammatory process going on in the bile ducts that damages them. They become narrow and blocked by scar tissue.
PSC is estimated to affect roughly 1 in 100,000 people yearly, and it is more prevalent in northern regions and among non-smokers.[ref] Since most patients will require a liver transplant, the survival rate historically has been only 12- 17 years after a diagnosis.[ref][ref]
Many people with PSC have coexisting autoimmune conditions. Inflammatory bowel diseases (IBD), such as ulcerative colitis or Crohn’s disease, are reported in about 2/3 of PSC patients. About 25% of PSC patients also have other autoimmune diseases such as type 1 diabetes, rheumatoid arthritis, autoimmune thyroid disease, or psoriasis.[ref] These coexisting conditions may help shed some light on why PSC occurs.
What are the symptoms of PSC?
The early symptoms of PSC include:[ref]
- Fatigue
- Itching
- Jaundice ( yellowing of the eyes/skin)
- Abdominal pain
Often, this disease is diagnosed with routine blood tests (such as elevated levels of alkaline phosphatase) and magnetic resonance imaging of the liver before symptoms appear. Those with an early diagnosis may still feel well for years. Since no cause has been established, doctors are unclear on how this disease will progress for each individual.[ref]
However, as the disease progresses, the symptoms will run the range of a diseased liver. A person can expect to see a fever, chills, night sweats, enlarged liver, enlarged spleen, and weight loss.
Are there any complications to this disease?
Complications of PSC can include bleeding of the veins in the GI tract (variceal hemorrhage), cancer of the bile ducts, colon cancer, or gallbladder cancer.
Numerous novel therapies have been tested in PSC, although there is little proof that they are helpful. For end-stage liver disease brought on by PSC, liver transplantation effectively increases survival. After transplantation, however, PSC may return in about 15 – 20% of cases.
Cholangiocarcinoma (cancer of the bile ducts) is the PSC complication that is most feared. This extremely aggressive tumor is challenging to diagnose, and there are no indicators for early identification. Although liver transplantation with neoadjuvant chemo-irradiation shows promise in certain individuals with cholangiocarcinoma, the results are incredibly dismal.[ref][ref]
Other risk factors:
- Patients with a coexisting IBD disease have an increased risk of colon cancer.[ref]
- There is a chance of developing autoimmune conditions, like autoimmune hepatitis.[ref]
- Other autoimmune diseases that present alongside PSC include sarcoidosis, thyroid disease, and type I diabetes mellitus.[ref]
Theories on the Causes of PSC:
Researchers have several theories as to how and why PSC develops.
1) Aberrant homing theory:
T cells, which are part of our immune system, are thought to be misdirected from the gut to the liver due to altered receptor expression in the liver. This misdirection is referred to as aberrant homing. MadCAM-1 and CCL25 appear to be important actors. MadCAM-1 and CCL25, normally only expressed in the gut, are expressed in the livers of PSC patients leading to inflammation.[ref][ref]
A new study, though, shows that the aberrant T cells are found in all types of liver disease, not just PSC. The authors of the study contend that this new research negates the theory that aberrant T cells are causal for PSC.[ref]
2) Autoantibodies:
Antibodies against proteins in the body can cause the immune system to attack your own normal cells. One PSC theory is that antibodies drive an immune response that causes liver damage.
There seems to be a substantial genetic link between PSC and HLA (human leukocyte antigens). Different HLA types are strongly linked to various autoimmune diseases. Twenty-five years ago, researchers discovered a genetic link between PSC and the human leukocyte antigen (HLA) complex on chromosome 6p21.[ref]
For PSC, the association with pANCA antibodies (antineutrophil cytoplasmic antibodies that target a protein called MPO) supports the “autoimmunity” theory.[ref] In addition to pANCA antibodies, autoantibodies to Saccharomyces cerevisiae are also commonly detected. Saccharomyces cerevisiae is a common yeast species used in winemaking and baking. This yeast species is also a common finding in people with IBD.[ref] Another antibody found in IBD and PSC patients is anti-GP2. Glycoprotein 2 (GP2) is secreted from the pancreas along with digestive enzymes.[ref][ref]
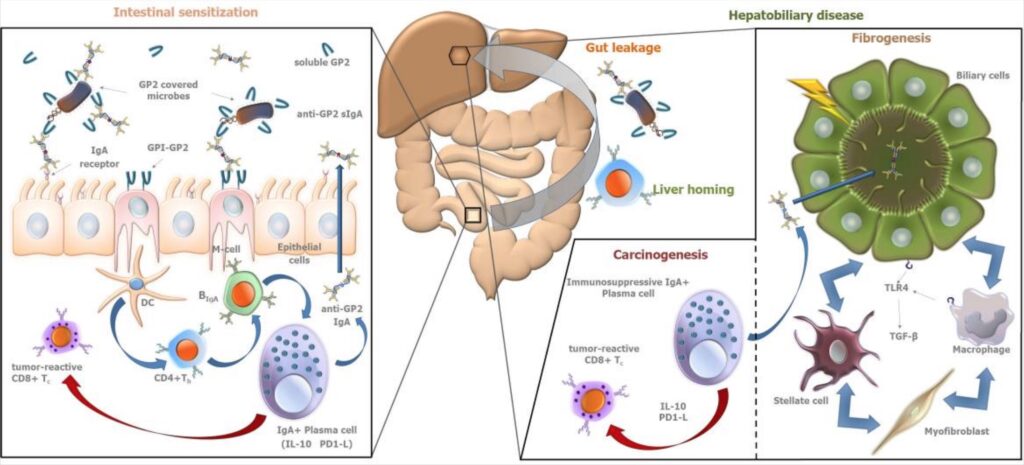
3) Bile Transporter Problems: Toxic bile theory
The “toxic bile” theory is based on the idea that bile transporter abnormalities, such as those in CFTR (cystic fibrosis gene) and ABCB4, could cause bile ducts to be chemically damaged by bile acids.[ref] CFTR is a protein that helps regulate salt and water on surfaces in the body, such as the lungs. Mutations in the CFTR gene can cause cystic fibrosis (check your CFTR gene here).[ref] ABCB4 codes for a protein that moves phospholipids across membranes in liver cells.[ref].
The steroid and xenobiotic receptor (SXR), also known as the pregnane X receptor (PXR), is a crucial nuclear receptor that controls bile acid detoxification and alternate excretion pathways. It has been demonstrated that PSC patients with certain genetic SXR/PXR variations experience a more severe illness trajectory. It is consistent with the significance of PXR’s protective actions in cholestasis-related animal models.
4) Leaky gut:
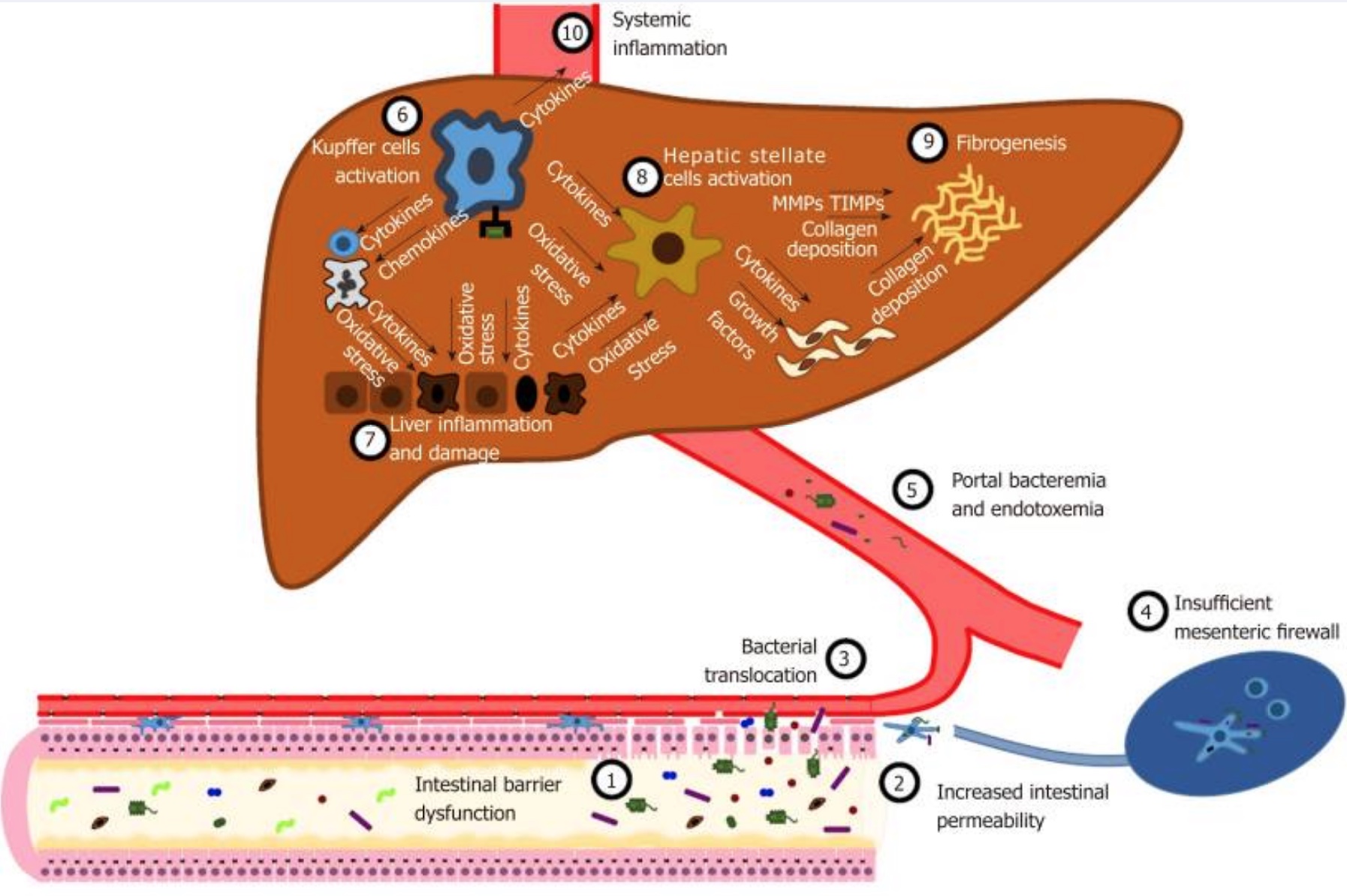
A leaky gut could be a factor as well. Leaky gut refers to the intestinal lining becoming more permeable, allowing gut microbes the ability to pass through. The microbes enter the bloodstream and end up in the liver. Early studies in a rat model of intestinal bacterial overgrowth with the “leaky gut” theory (conducted about 20 years ago) suggested that innate immune responses to bacterial metabolites may start a PSC-like pathogenesis.[ref]
While the liver is well-equipped to take care of some bacteria and bacterial components, a leaky gut and the translocation of bacteria to the liver is a driving factor in liver disease, such as NAFLD (non-alcoholic fatty liver disease).[ref]
5) Epstein-Barr Infection:
A new study suggests that Epstein-Barr infection may be linked to PSC. The study found that patients with PSC were more likely to have Epstein-Barr antibodies in their sampled T and B cells. The researchers also found that a mono infection increased the risk of PSC by 12-fold.[ref]
Related article: Epstein-Barr Virus
Primary Sclerosing Cholangitis Genotype Report:
Access this content:
An active subscription is required to access this content.
Lifehacks:
Primary sclerosing cholangitis is a serious, life-altering disease. You should discuss any dietary or supplement changes with your doctor.
The rest of this article is for Genetic Lifehacks members only. Consider joining today to see the rest of this article.
Access this content:
An active subscription is required to access this content.
Related Articles and Topics:
NAFLD Genes:
Fatty liver disease, NAFLD, is caused by a buildup of fat in the liver. Genetics plays a role – along with diet and environmental toxins.
Chronic Inflammation:
Most chronic diseases have inflammation as an underlying cause. Find out how your genes impact your risk of elevated inflammatory cytokines.
Curcumin Supplements: Decreasing Inflammation
Have you heard that curcumin supplements offer a slew of health benefits? Discover the science behind how curcumin reduces inflammation for better outcomes in chronic diseases.
Autoimmune Diseases: Genetics plus Triggers
A list of articles for an in-depth look at the background science, research studies, and genetic variants related to the triggers of autoimmune diseases.
References:
ABCB4 Gene: MedlinePlus Genetics. https://medlineplus.gov/genetics/gene/abcb4/. Accessed 11 Aug. 2022.
Adams, David H., and Bertus Eksteen. “Aberrant Homing of Mucosal T Cells and Extra-Intestinal Manifestations of Inflammatory Bowel Disease.” Nature Reviews. Immunology, vol. 6, no. 3, Mar. 2006, pp. 244–51. PubMed, https://doi.org/10.1038/nri1784.
Antineutrophil Cytoplasmic Antibodies (ANCA) Test: MedlinePlus Medical Test. https://medlineplus.gov/lab-tests/antineutrophil-cytoplasmic-antibodies-anca-test/. Accessed 11 Aug. 2022.
Bai, Xue, et al. “Effects of SLCO1B1 and GATM Gene Variants on Rosuvastatin-Induced Myopathy Are Unrelated to High Plasma Exposure of Rosuvastatin and Its Metabolites.” Acta Pharmacologica Sinica, vol. 40, no. 4, Apr. 2019, pp. 492–99. PubMed, https://doi.org/10.1038/s41401-018-0013-y.
Carr, Daniel F., et al. “Genomewide Association Study of Statin-Induced Myopathy in Patients Recruited Using the UK Clinical Practice Research Datalink.” Clinical Pharmacology and Therapeutics, vol. 106, no. 6, Dec. 2019, pp. 1353–61. PubMed, https://doi.org/10.1002/cpt.1557.
Chung, Sharon A., et al. “Differential Genetic Associations for Systemic Lupus Erythematosus Based on Anti–DsDNA Autoantibody Production.” PLoS Genetics, vol. 7, no. 3, Mar. 2011, p. e1001323. PubMed Central, https://doi.org/10.1371/journal.pgen.1001323.
Fernando, Michelle M. A., et al. “Identification of Two Independent Risk Factors for Lupus within the MHC in United Kingdom Families.” PLoS Genetics, vol. 3, no. 11, Nov. 2007, p. e192. PubMed Central, https://doi.org/10.1371/journal.pgen.0030192.
Ferri, Priscila Menezes, et al. “The Role of Genetic and Immune Factors for the Pathogenesis of Primary Sclerosing Cholangitis in Childhood.” Gastroenterology Research and Practice, vol. 2016, 2016, p. 3905240. PubMed Central, https://doi.org/10.1155/2016/3905240.
Fickert, Peter, et al. “Regurgitation of Bile Acids from Leaky Bile Ducts Causes Sclerosing Cholangitis in Mdr2 (Abcb4) Knockout Mice.” Gastroenterology, vol. 127, no. 1, July 2004, pp. 261–74. PubMed, https://doi.org/10.1053/j.gastro.2004.04.009.
Folseraas, Trine, et al. “Extended Analysis of a Genome-Wide Association Study in Primary Sclerosing Cholangitis Detects Multiple Novel Risk Loci.” Journal of Hepatology, vol. 57, no. 2, Aug. 2012, pp. 366–75. PubMed Central, https://doi.org/10.1016/j.jhep.2012.03.031.
Graham, Jonathon J., et al. “Aberrant Hepatic Trafficking of Gut-Derived T-Cells Is Not Specific to Primary Sclerosing Cholangitis.” Hepatology (Baltimore, Md.), vol. 75, no. 3, Mar. 2022, pp. 518–30. PubMed Central, https://doi.org/10.1002/hep.32193.
Hirschfield, Gideon M., et al. “Primary Sclerosing Cholangitis.” Lancet (London, England), vol. 382, no. 9904, Nov. 2013, pp. 1587–99. PubMed, https://doi.org/10.1016/S0140-6736(13)60096-3.
Karlsen, Tom H, et al. “Genetic Epidemiology of Primary Sclerosing Cholangitis.” World Journal of Gastroenterology : WJG, vol. 13, no. 41, Nov. 2007, pp. 5421–31. PubMed Central, https://doi.org/10.3748/wjg.v13.i41.5421.
Karlsen, Tom H., Andre Franke, et al. “Genome-Wide Association Analysis in Primary Sclerosing Cholangitis.” Gastroenterology, vol. 138, no. 3, Mar. 2010, pp. 1102–11. PubMed, https://doi.org/10.1053/j.gastro.2009.11.046.
Karlsen, Tom H., Trine Folseraas, et al. “Primary Sclerosing Cholangitis – a Comprehensive Review.” Journal of Hepatology, vol. 67, no. 6, Dec. 2017, pp. 1298–323. www.journal-of-hepatology.eu, https://doi.org/10.1016/j.jhep.2017.07.022.
Kotronen, A., et al. “A Common Variant in PNPLA3, Which Encodes Adiponutrin, Is Associated with Liver Fat Content in Humans.” Diabetologia, vol. 52, no. 6, June 2009, pp. 1056–60. PubMed, https://doi.org/10.1007/s00125-009-1285-z.
Lamberts, Laetitia E., et al. “Immune-Mediated Diseases in Primary Sclerosing Cholangitis.” Digestive and Liver Disease: Official Journal of the Italian Society of Gastroenterology and the Italian Association for the Study of the Liver, vol. 43, no. 10, Oct. 2011, pp. 802–06. PubMed, https://doi.org/10.1016/j.dld.2011.05.009.
LaRusso, Nicholas F., et al. “Primary Sclerosing Cholangitis: Summary of a Workshop.” Hepatology (Baltimore, Md.), vol. 44, no. 3, Sept. 2006, pp. 746–64. PubMed, https://doi.org/10.1002/hep.21337.
Lee, Hannah H., and Richard H. Ho. “Interindividual and Interethnic Variability in Drug Disposition: Polymorphisms in Organic Anion Transporting Polypeptide 1B1 (OATP1B1; SLCO1B1 ): OATP Polymorphisms and Drug Disposition.” British Journal of Clinical Pharmacology, vol. 83, no. 6, June 2017, pp. 1176–84. DOI.org (Crossref), https://doi.org/10.1111/bcp.13207.
Li, Xingang, et al. “The Association between Genetic Polymorphisms and Simvastatin-Induced Myopathy: A Narrative Synthesis of Evidence.” Drug Research, vol. 69, no. 4, Apr. 2019, pp. 185–93. PubMed, https://doi.org/10.1055/a-0677-2944.
Lichtman, S. N., et al. “Biliary Tract Disease in Rats with Experimental Small Bowel Bacterial Overgrowth.” Hepatology (Baltimore, Md.), vol. 13, no. 4, Apr. 1991, pp. 766–72.
Macaluso, Fabio Salvatore, et al. “Genetic Background in Nonalcoholic Fatty Liver Disease: A Comprehensive Review.” World Journal of Gastroenterology : WJG, vol. 21, no. 39, Oct. 2015, pp. 11088–111. PubMed Central, https://doi.org/10.3748/wjg.v21.i39.11088.
Manganis, Charis D., et al. “Review of Primary Sclerosing Cholangitis with Increased IgG4 Levels.” World Journal of Gastroenterology, vol. 26, no. 23, June 2020, pp. 3126–44. PubMed Central, https://doi.org/10.3748/wjg.v26.i23.3126.
Mitchell, S. A., et al. “Association of the Tumour Necrosis Factor Alpha -308 but Not the Interleukin 10 -627 Promoter Polymorphism with Genetic Susceptibility to Primary Sclerosing Cholangitis.” Gut, vol. 49, no. 2, Aug. 2001, pp. 288–94. PubMed, https://doi.org/10.1136/gut.49.2.288.
Nicoletti, Alberto, et al. “Intestinal Permeability in the Pathogenesis of Liver Damage: From Non-Alcoholic Fatty Liver Disease to Liver Transplantation.” World Journal of Gastroenterology, vol. 25, no. 33, Sept. 2019, pp. 4814–34. PubMed Central, https://doi.org/10.3748/wjg.v25.i33.4814.
Primary Sclerosing Cholangitis – Symptoms and Causes. https://www.mayoclinic.org/diseases-conditions/primary-sclerosing-cholangitis/symptoms-causes/syc-20355797. Accessed 11 Aug. 2022.
Rupp, C., et al. “Fut2 Genotype Is a Risk Factor for Dominant Stenosis and Biliary Candida Infections in Primary Sclerosing Cholangitis.” Alimentary Pharmacology & Therapeutics, vol. 39, no. 8, Apr. 2014, pp. 873–82. PubMed, https://doi.org/10.1111/apt.12663.
“SLCO1B1 Variants and Statin-Induced Myopathy — A Genomewide Study.” New England Journal of Medicine, vol. 359, no. 8, Aug. 2008, pp. 789–99. nejm.org (Atypon), https://doi.org/10.1056/NEJMoa0801936.
Tornai, David, et al. “Serological Biomarkers for Management of Primary Sclerosing Cholangitis.” World Journal of Gastroenterology, vol. 28, no. 21, June 2022, pp. 2291–301. PubMed Central, https://doi.org/10.3748/wjg.v28.i21.2291.
Tornai, David, and Maria Papp. “Editorial: Serologic Antibodies in Primary Sclerosing Cholangitis – a Tell‐tale Sign of Compromised Gut‐liver Immunity?” Alimentary Pharmacology & Therapeutics, vol. 53, no. 2, Jan. 2021, pp. 350–51. DOI.org (Crossref), https://doi.org/10.1111/apt.16201.
Trépo, Eric, et al. “Impact of Patatin-like Phospholipase-3 (Rs738409 C>G) Polymorphism on Fibrosis Progression and Steatosis in Chronic Hepatitis C.” Hepatology (Baltimore, Md.), vol. 54, no. 1, July 2011, pp. 60–69. PubMed, https://doi.org/10.1002/hep.24350.
Wijarnpreecha, Karn, et al. “Association between Smoking and Risk of Primary Sclerosing Cholangitis: A Systematic Review and Meta-Analysis.” United European Gastroenterology Journal, vol. 6, no. 4, May 2018, pp. 500–08. PubMed, https://doi.org/10.1177/2050640618761703.