Key takeaways:
~ Hemochromatosis is a relatively common genetic disease that causes iron to build up in the body, damaging organs and cells, which leads to chronic diseases and eventually death.
~ Knowing that you carry the genetic variants for hemochromatosis means that you can prevent iron buildup by giving blood.
~ 23andMe and AncestryDNA genetic data can tell you if you likely carry the more common genetic mutations in the HFE gene for hemochromatosis.
What is hemochromatosis?
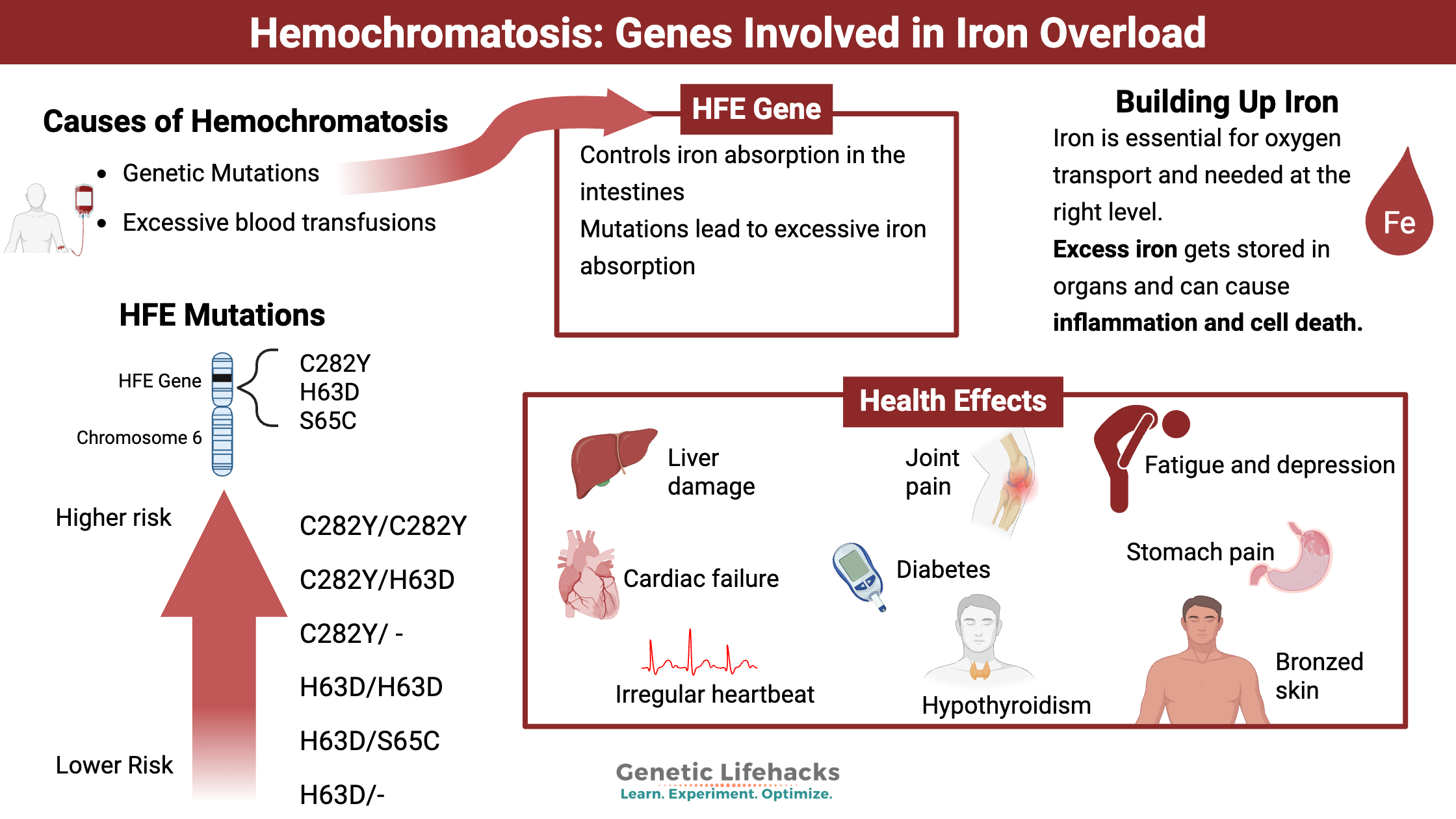
Hemochromatosis is a disease state of building up too much iron in the body. It can be caused by genetic mutations in iron-related genes, or it can be due to excessive blood transfusions for anemia or liver disease.
Iron is essential – but it needs to be in just the right amount. People with hereditary hemochromatosis (also spelled haemochromatosis) absorb more iron from food than they should due to genetic variants.
Once iron gets absorbed in the intestines, the body doesn’t have a good way to get rid of it. Usually, the body tightly regulates the amount of iron absorbed. When too much iron is absorbed, such as for people carrying genetic variants that cause hemochromatosis, the excess builds up over time. The body stores the excess iron in the organs, including the liver, heart, pancreas, skin, pituitary, and thyroid. You can also store iron in your joints and bone marrow.[ref]
What are the symptoms of hemochromatosis?
Symptoms of building up too much iron can include:
- joint pain
- tiredness
- abdominal pain
- liver disease
- diabetes
- irregular heartbeat
- bronzed skin color
- hypothyroidism
- elevated blood sugar, and more.
Why do we need iron?
The body uses iron in numerous ways. In fact, every cell in the body uses iron in one form or another. Iron, involved in redox reactions, moves electrons in the mitochondria to produce energy. Also, red blood cells need iron as a critical component for moving oxygen throughout the body.
Rusty old iron parts may come to mind when you think about iron out in nature. Rust forms because iron can be easily oxidized, forming compounds with oxygen. This ability to move electrons around makes iron incredibly important and incredibly reactive in the body.
Regulating Iron in the Body:
The body regulates the absorption of iron tightly. We do not want too much oxidation occurring in the wrong place (no ‘rusting’ in the body ;-), but our body also regulates iron to keep it away from iron-loving bacteria. Certain pathogenic species of bacteria thrive and multiply rapidly when iron is available. Thus, we don’t want too much iron in the body, and we have ways of quickly sequestering, or storing, the iron when the body is fighting off a pathogen.[ref]
The liver produces the iron regulatory hormone called hepcidin. Discovered and named in 2000, scientists have since figured out that hepcidin controls the regulation of iron in the body and responds to lipopolysaccharides, a component of bacteria, to prevent iron-loving bacteria from reproducing rapidly.[ref]
The HFE gene was discovered in 1996 and is linked to hereditary hemochromatosis. This gene codes for a protein that controls how iron is taken into the intestinal cells. Free iron is bound to transferrin, which binds to a transferrin receptor to be taken into the cell. HFE can also bind to the transferrin receptor, blocking the receptor from taking iron into the cells. In this way, HFE is a negative regulator of iron absorption.[ref] As an iron sensor, HFE controls iron intake and regulates the downstream production of hepcidin.[ref]
When too much iron is absorbed, whether because of a mutation in the HFE gene or other genes regulating iron, the body stores it in different organs. Excess iron causes oxidative stress and DNA damage to the tissue. This excess can cause fibrosis and cell death, leading to liver cirrhosis or other organ damage.[ref]
How do you inherit hemochromatosis?
There are multiple causes of hereditary hemochromatosis. The most common cause is a genetic variant that impacts the HFE gene. It allows too much iron to be absorbed into the body and is called type 1, or classic hemochromatosis.
Other rare causes of hemochromatosis include variations in other genes involved in hepcidin regulation and iron transport. These are known as hemochromatosis types 2, 3, and 4.
Interestingly, hemochromatosis was first described in the 1700s as ‘bronze diabetes’ or ‘pigmented cirrhosis. It refers to how iron accumulates in the skin and can make it look bronzed, which combines with the higher risk of diabetes or cirrhosis in hemochromatosis.[ref]
How rare is hemochromatosis?
The HFE mutations are much more common in people of Northern European heritage, but not everyone who carries the variants will end up with a diagnosis of hemochromatosis.
Estimates show that ~10% of people of Northern European ancestry carry one copy of an HFE variant (more on this below). It is thought that the primary hemochromatosis mutation (HFE C282Y) originated in the Celtic or Viking population before 4,000 BC. From there, it spread through the European population.[ref]
Studies on hemochromatosis vary quite a bit in their estimates of how common hemochromatosis is among people with the variants, mainly due to differences in the clinical definition.
Some define hemochromatosis as having a ferritin (a measure of storage iron) level of over 1,000 mcg/l, liver disease, heart failure, and bronzed skin. Others define hemochromatosis as having elevated ferritin (>300 mcg/l) and liver failure.[ref]
The question, then, is at which point in the spectrum of organ damage do physicians define hemochromatosis – is just having the pancreas damaged and causing diabetes enough, or do you also need liver damage? Clinical diagnosis and research study definitions vary quite a bit.
But this website is all about optimizing health, so let’s look at what happens to the body before your liver fails…
Problems with excess iron well before a hemochromatosis diagnosis:
As you can imagine, the symptoms of iron building up can occur over the decades leading up to the point of organ failure.
Symptoms of excess iron include:
- Joint pain, fatigue, and irritability are common complaints in people with HFE genetic variants.
- Fibromyalgia and arthritis are common diagnoses.[ref][ref]
- Reduced libido and erectile dysfunction in men are common with iron buildup. [ref]
- Diabetes and heart disease diagnoses can often precede the clinical diagnosis of hemochromatosis.[ref]
The key here is that all health problems related to excess iron can be prevented.
You aren’t destined to have aches and pains leading up to liver failure or diabetes — knowledge is power here, and iron levels can be reduced. (More on this in the Lifehacks section below).
Carrying just one copy of the HFE variant (mainly C282Y):
The initial issues with too much iron include random joint pain, fatigue, irritability, and/or abdominal pain. Research studies also show that carriers of HFE variants (mainly C282Y) are at a higher risk of :
- non-alcoholic fatty liver disease[ref][ref]
- metabolic syndrome[ref]
- cardiovascular disease[ref] including women with heterozygous variants[ref]
- slightly higher risk of cancer[ref][ref] including breast cancer[ref] and liver cancer[ref]
- Alzheimer’s disease, including heterozygous carriers[ref]
- musculoskeletal problems (arthritis symptoms)[ref]
- high blood pressure[ref]
- high uric acid (gout[ref]
- lung fibrosis[ref]
- diabetes[ref]
- cardiovascular disease in kidney disease patients[ref]
- increased lead levels[ref][ref]
HFE and the Immune System:
While iron is essential for transporting oxygen throughout the body, it can also be essential for bacterial growth. As I mentioned above, many bacterial species thrive in iron-rich environments. One major role of iron regulation in the body is iron storage via ferritin to sequester iron away from bacteria.[ref] A 2024 study of over 140,000 Danish people found that those who carried the C282Y variant associated with hemochromatosis were at an increased risk of infection, sepsis, and death from infection. The study noted that even those who carried the C282Y variant but had normal iron levels were at an increased risk of infection. [ref]
While iron regulation and the HFE gene are important in bacterial infections, it turns out that HFE may play a role in the immune system that is not related to iron. In fact, researchers initially classified the HFE gene as part of the HLA system[ref]
Research shows that a protein in HIV significantly downregulates the HFE (iron homeostasis) gene.[ref] Researchers have also found that infection with cytomegalovirus decreases HFE by degrading the protein.
The question for researchers has been, ‘How does interfering with iron and downregulating the HFE protein help viruses reproduce?’.
It turns out that the answer may be that HFE plays a role in the innate immune response that is separate from iron homeostasis. Recent research shows that irrespective of iron levels, HFE interacts with interferon, which is the body’s initial fighter against viral infections. HFE deficiency (in mice) increased interferon and made the animals more resistant to the flu virus.[ref]
Excess iron also interferes with active T cell function by increasing the checkpoints that decrease T cell function and cause exhaustion.[ref]
Related article: T cell exhaustion
Benefits of an HFE mutation? Tradeoffs.
The negative impacts of too much iron are obvious, but it is likely that the mutations are relatively common in the population due to a survival benefit.
- Women with higher iron levels would be less likely to die in childbirth and thus pass on the mutation to their children.
- A 2021 study found a benefit for athletes with the HFE mutation. Competitive cyclists with HFE mutations have faster performance times and a 17% greater V˙O2 peak.[ref]
- A 2020 study found that international-level endurance athletes were almost twice as likely to carry the HFE H63D variant. The researchers think this is due to increased oxygen-binding capacity with higher iron.[ref]
Hemochromatosis Genotype Report:
HFE Gene Mutations:
Access this content:
An active subscription is required to access this content.
Lifehacks:
Blood tests:
Adult men and menopausal women who are heterozygous or homozygous for any of the HFE variants should, in my not-a-doctor opinion, get their serum iron, TIBC, and ferritin levels checked or ask their doctor to test them.
Ordering serum iron w/ TIBC and ferritin should give you enough information to know if you are storing too much iron. If you want to order your own test, an inexpensive option in the US is UltaLabs. If you wish your insurance or country’s health services to cover the test, talk with your doctor and explain that you carry a genetic mutation in the HFE gene.
Give Blood Regularly
If you have slightly elevated iron levels, the simplest way to manage them is to give blood regularly. You will likely feel good, and you will definitely help out someone else with your blood donation. It is a win-win.
Fatty liver disease risk:
People with HFE mutations and hemochromatosis are at an increased risk of fatty liver disease (NAFLD). Talk with your doctor about testing for NAFLD.
Related article: Fatty Liver Disease: Genetic variants that increase the risk of NAFLD
Natural Iron Chelators and Inhibitors:
Access this content:
An active subscription is required to access this content.
Related Articles and Topics:
Lithium Orotate and Vitamin B12: Benefits for Mood and Cognitive Support
Is Inflammation Causing Your Depression and Anxiety? The Science Behind the Link
Alzheimer’s Gene: Find your APOE type from your genetic raw data
References:
Ali-Rahmani, Fatima, et al. “HFE Gene Variants, Iron, and Lipids: A Novel Connection in Alzheimer’s Disease.” Frontiers in Pharmacology, vol. 5, July 2014, p. 165. PubMed Central, https://doi.org/10.3389/fphar.2014.00165.
Basuli, Debargha, et al. “Epidemiological Associations between Iron and Cardiovascular Disease and Diabetes.” Frontiers in Pharmacology, vol. 5, May 2014, p. 117. PubMed Central, https://doi.org/10.3389/fphar.2014.00117.
Ben-Arieh, Sayeh Vahdati, et al. “Human Cytomegalovirus Protein US2 Interferes with the Expression of Human HFE, a Nonclassical Class I Major Histocompatibility Complex Molecule That Regulates Iron Homeostasis.” Journal of Virology, vol. 75, no. 21, Nov. 2001, pp. 10557–62. PubMed Central, https://doi.org/10.1128/JVI.75.21.10557-10562.2001.
Camacho, António, et al. “Effect of C282Y Genotype on Self-Reported Musculoskeletal Complications in Hereditary Hemochromatosis.” PloS One, vol. 10, no. 3, 2015, p. e0122817. PubMed, https://doi.org/10.1371/journal.pone.0122817.
Castiella, Agustin, et al. “Impact of H63D Mutations, Magnetic Resonance and Metabolic Syndrome among Outpatient Referrals for Elevated Serum Ferritin in the Basque Country.” Annals of Hepatology, vol. 14, no. 3, 2015, pp. 333–39.
Distante, S., et al. “The Origin and Spread of the HFE-C282Y Haemochromatosis Mutation.” Human Genetics, vol. 115, no. 4, Sept. 2004, pp. 269–79. PubMed, https://doi.org/10.1007/s00439-004-1152-4.
Drakesmith, Hal, et al. “HIV-1 Nef down-Regulates the Hemochromatosis Protein HFE, Manipulating Cellular Iron Homeostasis.” Proceedings of the National Academy of Sciences of the United States of America, vol. 102, no. 31, Aug. 2005, pp. 11017–22. PubMed, https://doi.org/10.1073/pnas.0504823102.
Ellervik, Christina, et al. “Total Mortality by Elevated Transferrin Saturation in Patients With Diabetes.” Diabetes Care, vol. 36, no. 9, Sept. 2013, pp. 2646–54. PubMed Central, https://doi.org/10.2337/dc12-2032.
Flais, Jérémy, et al. “Hyperferritinemia Increases the Risk of Hyperuricemia in HFE-Hereditary Hemochromatosis.” Joint Bone Spine, vol. 84, no. 3, May 2017, pp. 293–97. PubMed, https://doi.org/10.1016/j.jbspin.2016.05.020.
Grosse, Scott D., et al. “Clinical Penetrance in Hereditary Hemochromatosis: Estimates of the Cumulative Incidence of Severe Liver Disease among HFE C282Y Homozygotes.” Genetics in Medicine, vol. 20, no. 4, Apr. 2018, pp. 383–89. www.nature.com, https://doi.org/10.1038/gim.2017.121.
Hollerer, Ina, et al. “Pathophysiological Consequences and Benefits of HFE Mutations: 20 Years of Research.” Haematologica, vol. 102, no. 5, May 2017, pp. 809–17. haematologica.org, https://doi.org/10.3324/haematol.2016.160432.
Kayaalti, Zeliha, et al. “Maternal Hemochromatosis Gene H63D Single-Nucleotide Polymorphism and Lead Levels of Placental Tissue, Maternal and Umbilical Cord Blood.” Environmental Research, vol. 140, July 2015, pp. 456–61. PubMed, https://doi.org/10.1016/j.envres.2015.05.004.
Kemna, Erwin H. J. M., et al. “Hepcidin: From Discovery to Differential Diagnosis.” Haematologica, vol. 93, no. 1, Jan. 2008, pp. 90–97. PubMed, https://doi.org/10.3324/haematol.11705.
Liu, Juan, et al. “HFE Inhibits Type I IFNs Signaling by Targeting the SQSTM1-Mediated MAVS Autophagic Degradation.” Autophagy, vol. 17, no. 8, pp. 1962–77. PubMed Central, https://doi.org/10.1080/15548627.2020.1804683. Accessed 10 Feb. 2026.
Liu, Xiaoyan, et al. “C282Y Polymorphism in the HFE Gene Is Associated with Risk of Breast Cancer.” Tumour Biology: The Journal of the International Society for Oncodevelopmental Biology and Medicine, vol. 34, no. 5, Oct. 2013, pp. 2759–64. PubMed, https://doi.org/10.1007/s13277-013-0833-9.
Lopez, Christopher A., and Eric P. Skaar. “The Impact of Dietary Transition Metals on Host-Bacterial Interactions.” Cell Host & Microbe, vol. 23, no. 6, June 2018, pp. 737–48. PubMed Central, https://doi.org/10.1016/j.chom.2018.05.008.
Määttä, Kirsi M., et al. “Genetic Variant Coding for Iron Regulatory Protein HFE Contributes to Hypertension, the TAMRISK Study.” Medicine, vol. 94, no. 4, Jan. 2015, p. e464. PubMed, https://doi.org/10.1097/MD.0000000000000464.
Milman, Nils Thorm, et al. “Diagnosis and Treatment of Genetic HFE-Hemochromatosis: The Danish Aspect.” Gastroenterology Research, vol. 12, no. 5, Oct. 2019, pp. 221–32. PubMed Central, https://doi.org/10.14740/gr1206.
———. “Diagnosis and Treatment of Genetic HFE-Hemochromatosis: The Danish Aspect.” Gastroenterology Research, vol. 12, no. 5, Oct. 2019, pp. 221–32. PubMed, https://doi.org/10.14740/gr1206.
Mohammad, Ausaf, et al. “High Prevalence of Fibromyalgia in Patients with HFE-Related Hereditary Hemochromatosis.” Journal of Clinical Gastroenterology, vol. 47, no. 6, July 2013, pp. 559–64. PubMed, https://doi.org/10.1097/MCG.0b013e31826f7ad7.
Nigg, Joel T., et al. “Variation in an Iron Metabolism Gene Moderates the Association Between Blood
Lead Levels and Attention-Deficit/Hyperactivity Disorder in Children.” Psychological Science, vol. 27, no. 2, Feb. 2016, pp. 257–69. PubMed Central, https://doi.org/10.1177/0956797615618365.
Page, Malcom G. P. “The Role of Iron and Siderophores in Infection, and the Development of Siderophore Antibiotics.” Clinical Infectious Diseases: An Official Publication of the Infectious Diseases Society of America, vol. 69, no. Suppl 7, Dec. 2019, pp. S529–37. PubMed Central, https://doi.org/10.1093/cid/ciz825.
Reuben, Alexandre, et al. “The Hemochromatosis Protein HFE 20 Years Later: An Emerging Role in Antigen Presentation and in the Immune System.” Immunity, Inflammation and Disease, vol. 5, no. 3, Apr. 2017, pp. 218–32. PubMed Central, https://doi.org/10.1002/iid3.158.
Roest, M., et al. “Heterozygosity for a Hereditary Hemochromatosis Gene Is Associated with Cardiovascular Death in Women.” Circulation, vol. 100, no. 12, Sept. 1999, pp. 1268–73. PubMed, https://doi.org/10.1161/01.cir.100.12.1268.
Sangiuolo, Federica, et al. “HFE Gene Variants and Iron-Induced Oxygen Radical Generation in Idiopathic Pulmonary Fibrosis.” The European Respiratory Journal, vol. 45, no. 2, Feb. 2015, pp. 483–90. PubMed, https://doi.org/10.1183/09031936.00104814.
Semenova, Ekaterina A., et al. “The Association of HFE Gene H63D Polymorphism with Endurance Athlete Status and Aerobic Capacity: Novel Findings and a Meta-Analysis.” European Journal of Applied Physiology, vol. 120, no. 3, 2020, pp. 665–73. PubMed Central, https://doi.org/10.1007/s00421-020-04306-8.
Shen, L. L., et al. “Implicating the H63D Polymorphism in the HFE Gene in Increased Incidence of Solid Cancers: A Meta-Analysis.” Genetics and Molecular Research: GMR, vol. 14, no. 4, Oct. 2015, pp. 13735–45. PubMed, https://doi.org/10.4238/2015.October.28.36.
Thakkar, Drishti, et al. “HFE Genotype and Endurance Performance in Competitive Male Athletes.” Medicine and Science in Sports and Exercise, vol. 53, no. 7, July 2021, pp. 1385–90. PubMed, https://doi.org/10.1249/MSS.0000000000002595.
Ye, Qing, et al. “Association between the HFE C282Y, H63D Polymorphisms and the Risks of Non-Alcoholic Fatty Liver Disease, Liver Cirrhosis and Hepatocellular Carcinoma: An Updated Systematic Review and Meta-Analysis of 5,758 Cases and 14,741 Controls.” PloS One, vol. 11, no. 9, 2016, p. e0163423. PubMed, https://doi.org/10.1371/journal.pone.0163423.
Zhang, Meng, et al. “Meta-Analysis of the Association between H63D and C282Y Polymorphisms in HFE and Cancer Risk.” Asian Pacific Journal of Cancer Prevention: APJCP, vol. 16, no. 11, 2015, pp. 4633–39. PubMed, https://doi.org/10.7314/apjcp.2015.16.11.4633.
http://www.irondisorders.org/iron-overload. Accessed 10 Feb. 2026.
ZHANG, ZEYU, et al. “Taurine Supplementation Reduces Oxidative Stress and Protects the Liver in an Iron-Overload Murine Model.” Molecular Medicine Reports, vol. 10, no. 5, Nov. 2014, pp. 2255–62. PubMed Central, https://doi.org/10.3892/mmr.2014.2544.